Difference between revisions of "2022 DOCK tutorial 3 with PDBID 1X70"
BrockBoysan (talk | contribs) (→Receptor Prep) |
BrockBoysan (talk | contribs) (→Receptor Prep) |
||
| Line 40: | Line 40: | ||
Now that we the receptor by itself, we have to clean up the rest of the receptor by adding any missing side chains, dealing with multiple occupancies and mutated residues, and protonating and calculating partial charges. | Now that we the receptor by itself, we have to clean up the rest of the receptor by adding any missing side chains, dealing with multiple occupancies and mutated residues, and protonating and calculating partial charges. | ||
| − | For 1X70, there are several residues with multiple occupancies | + | For 1X70, there are several residues with multiple occupancies. |
| − | + | One way to check for these residues is to grep the number of alpha carbons from the pdb in the command line. | |
| − | + | In the terminal, type: grep -e CA 1X70_noH.pdb | |
[[File:1X70_multiple_occupancy.png|thumb|center|800px|Two potential occupancies shown for CYS649]] | [[File:1X70_multiple_occupancy.png|thumb|center|800px|Two potential occupancies shown for CYS649]] | ||
Revision as of 15:50, 1 March 2022
Contents
Introduction
DOCK
System Preparation
Fetching 1X70
Open Chimera and do the following to grab the protein:
File > Fetch By ID > 1X70
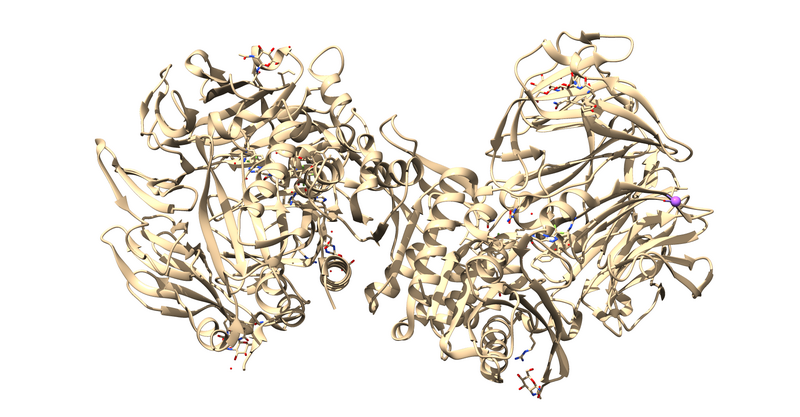
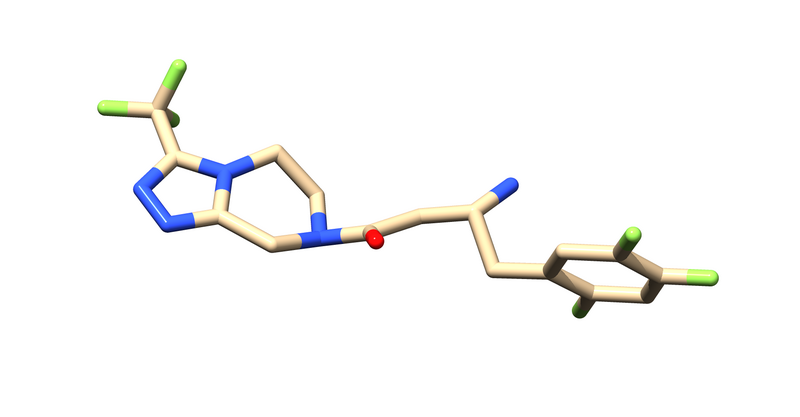
The first thing to notice is that this is a dimer and the ligand, 715, is not bound at the dimer interface. Thus, one of the monomers is entirely redundant and should be deleted.
Select > Chain > A
Actions > Atoms > Delete
Next it is important to remove cofactors, ions, and water molecules not involved in the binding interactions. This can be checked by reading through the paper associated with the PDB.
Select > Residue > NAG > Actions > Atoms > Delete
Select > Residue > NDG > Actions > Atoms > Delete
Select > Residue > HOH > Actions > Atoms > Delete
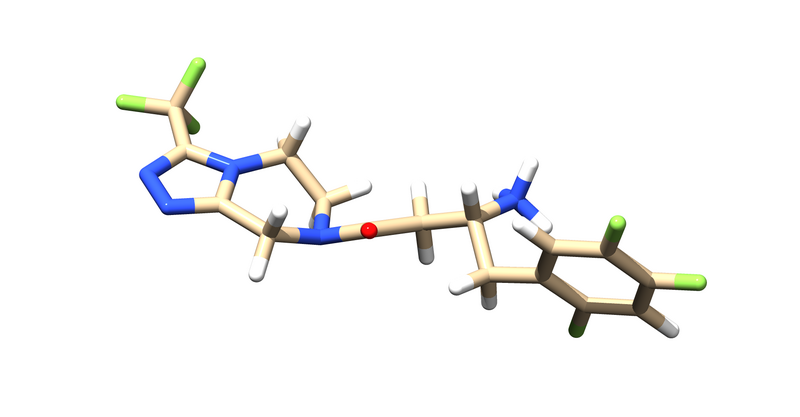
This will leave us with just the ligand and receptor.
Receptor Prep
Now that we the receptor by itself, we have to clean up the rest of the receptor by adding any missing side chains, dealing with multiple occupancies and mutated residues, and protonating and calculating partial charges.
For 1X70, there are several residues with multiple occupancies.
One way to check for these residues is to grep the number of alpha carbons from the pdb in the command line.
In the terminal, type: grep -e CA 1X70_noH.pdb
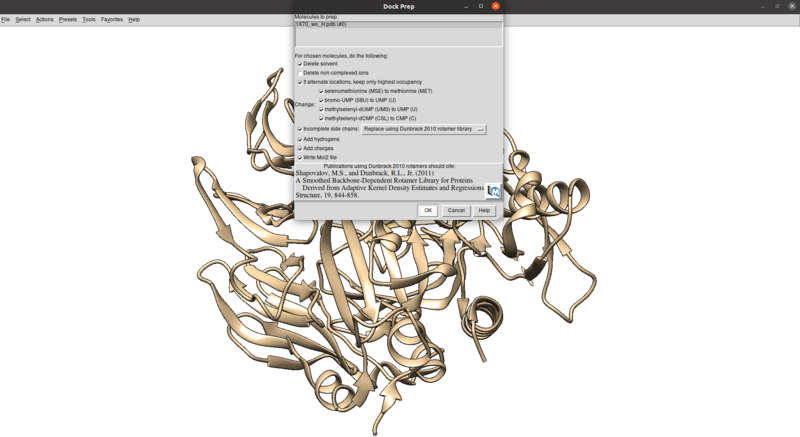
Tools > Structure Editing > Dock Prep
Make sure that the protonation makes sense for residues in the active site or coordinated with metals (none here), especially histidines, by checking the paper and making sure any nitrogens coordinating with metals are not protonated.
Ligand Prep
Surface Generation & Spheres
This section details the generation of sphere files which will be used to describe where you are trying to DOCK to on your protein.
Surface Generation In Chimera: Load 1X70 w/o Hydrogens > actions > show > surface
Then you will write a DMS (Molecular Surface) File With the surface generated in Chimera: Tools > Structure Editing > Write DMS
Now you should have a DMS file for the next step.
Sphere Generation To generate spheres make the following input file: "INSPH" -
1X70_dms.dms #Molecular Surface File R #Whether to generate spheres outside of surface (R) or inside (L) X #Surface points from the DMS file to use in sphere generation 0 #Minimum radius between spheres 4.0 #Maximum radius of sphere 1.4 #Minimum radius of sphere 1X70_wo_H.sph #Output sphere file
For more information on sphere generation see: https://dock.compbio.ucsf.edu/DOCK_6/tutorials/sphere_generation/generating_spheres.htm
The .sph file should give you something similar to the following image if you load it up over your protein in Chimera:
Sphere Selection
Making The Infamous Grid
Making the grid
Energy Minimization for the ligand
Docking & Virtual Screening
Rigid Docking
Fixed Anchor Docking
Flexible Docking
Placeholder
[[File:|thumb|center|800px|image placeholder]]