Difference between revisions of "2023 AMBER tutorial 1 with PDBID 4S0V"
Stonybrook (talk | contribs) (→Complex Equilibration) |
Stonybrook (talk | contribs) (→TLeap Implemenation) |
||
| Line 103: | Line 103: | ||
###write solvated toplogy and coordinate file | ###write solvated toplogy and coordinate file | ||
| − | saveamberparm solvcomplex 4s0v.wet.complex. | + | saveamberparm solvcomplex 4s0v.wet.complex.prmtop 4s0v.wet.complex.rst7 |
quit | quit | ||
Revision as of 07:43, 13 March 2023
Contents
Introduction
AMBER is a molecular dynamics program that can be run on your protein/ligand complex to ensure that the interactions between the two structures are stable. DOCK shows us how the two interact with each other at one point in time. AMBER looks at those interactions over time to ensure that forces will not occur which will push the ligand out of the binding site as the complex naturally moves. This tutorial will again be working with PDB #4s0v
Setting Up Your Environment
Just as with DOCK you should set up for directory structure at this point to keep everything organized and easy to find. We will be creating a new structure which looks like:
Structure
Before starting the analysis it's best to download a new protein/ligand complex from the PDB and isolate both the protein and ligand structures. Follow the steps in the *[1] tutorial to do this. The inputs we need are the isolated protein with NO hydrogens and NO charges; and the ligand with hydrogens and charges. In other words, once you isolate the protein structure in Chimera, save it with a filename such as, 4s0v_protein_for_AMBER.pdb. Then isolate the ligand structure, add hydrogens and re-do whatever protonation changes you made in the *[2] tutorial. Once the protonation state is correct, add charges and save the file as 4s0v_ligand_for_AMBER.mol2.
For the protein file, it’s important to model any missing loops before running AMBER on the complex. You might not have done this for the docking tutorial because the loops were far enough from the binding site to not matter but for this step it needs to be done.
Once these two files have been generated, scp them over to the 001.structure directory on Seawulf.
Force Field Parameters
AMBER needs force field parameters to run correctly. This only needs to be done for the ligand since information about the protein has been done over previous years. To generate the necessary files for AMBER we will be working on the command line on Seawulf, please cd into your 002.forceFieldParameters directory.
To generate the parameters for the ligand we need to run the following slurm script:
#!/bin/bash # #SBATCH --job-name=4s0v_AMBER_parameters #SBATCH --output=parameters_output.txt #SBATCH --ntasks-per-node=24 #SBATCH --nodes=6 #SBATCH --time=48:00:00 #SBATCH -p long-24core module load amber/16 antechamber -i ../001.structure/4s0v_ligand_for_AMBER.pdb -fi pdb -o 4s0v_ligand_antechamber.mol2 -fo mol2 -at gaff2 -c bcc -rn LIG -nc 0
The important parameter in the above command is the -nc option at the end. This is telling antechamber what the total charge is on your ligand. This number needs to be the same as the number used in the previous step when you added charges to the structure in Chimera.
Once this has completed running you will see multiple new files in your directory including, 4s0v_ligand_antechamber.mol2. This is the file with the parameters we just generated and what we need to use to generate the next file we need, a frcmod file, which will contain modified force field parameters. To generate the frcmod file, run the command:
parmchk2 -i 4s0v_ligand_antechamber.mol2 -f mol2 -o 4s0v_ligand.am1bcc.frcmod
If you get an error when running this line, try typing:
module load amber/16
and then running the command again. Once it's done running you will see the 4s0v_ligand.am1bcc.frcmod file in your directory.
TLeap Implemenation
Remembering that the point of running AMBER on our complex is to investigate the dynamics of the entire complex over time. In this next step we bring together the separate ligand and protein files so AMBER can be run on the whole comlex. This is done with a program called tleap. Again we will be working on the command line in Seawulf, please cd into your 003.tleap directory.
To create the input file:
vi 4s0v_tleap.in
and type the following lines into the input file:
#!/usr/bin/sh
###load protein force field
source leaprc.protein.ff14SB
###load GAFF force field (for our ligand)
source leaprc.gaff
###load TIP3P (water) force field
source leaprc.water.tip3p
###load ions frcmod for the tip3p model
loadamberparams frcmod.ionsjc_tip3p
###needed so we can use igb=8 model
set default PBradii mbondi3
###load protein pdb file
rec=loadpdb ../001.structure/4s0v_protein_for_AMBER.pdb
#bond rec.649.SG rec.762.SG
#bond rec.454.SG rec.472.SG
#bond rec.444.SG rec.447.SG
#bond rec.328.SG rec.339.SG
#bond rec.385.SG rec.394.SG
###load ligand frcmod/mol2
loadamberparams ../002.forceFieldParameters/4s0v_ligand.am1bcc.frcmod
lig=loadmol2 ../002.forceFieldParameters/4s0v_ligand_antechamber.mol2
###create gase-phase complex
gascomplex= combine {rec lig}
###write gas-phase pdb
savepdb gascomplex 4s0v.gas.complex.pdb
###write gase-phase toplogy and coord files for MMGBSA calc
saveamberparm gascomplex 4s0v.complex.parm7 4s0v.gas.complex.rst7
saveamberparm rec 4s0v.gas.receptor.parm7 4s0v.gas.receptor.rst7
saveamberparm lig 4s0v.gas.ligand.parm7 4s0v.gas.ligand.rst7
###create solvated complex (albeit redundant)
solvcomplex= combine {rec lig}
###solvate the system
solvateoct solvcomplex TIP3PBOX 12.0
###Neutralize system
addions solvcomplex Cl- 0
addions solvcomplex Na+ 0
#write solvated pdb file
savepdb solvcomplex 4s0v.wet.complex.pdb
###check the system
charge solvcomplex
check solvcomplex
###write solvated toplogy and coordinate file
saveamberparm solvcomplex 4s0v.wet.complex.prmtop 4s0v.wet.complex.rst7
quit
We should again run this input file using a slurm script:
#!/bin/bash # #SBATCH --job-name=4s0v_tleap #SBATCH --output=tleap_output.txt #SBATCH --ntasks-per-node=24 #SBATCH --nodes=6 #SBATCH --time=48:00:00 #SBATCH -p long-24core module load amber/16 tleap -f 4s0v_tleap.in
When this is done running you will see multiple files in your directory. The ones we will be working with are:
- 4s0v.wet.complex.parm7
- 4s0v.wet.complex.rst7
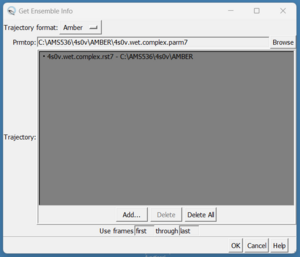
scp these over to your local computer and start a new session in Chimera. We need to load them by doing Tools → MD/Ensemble Analysis → MD Movie. A dialogue box will appear:
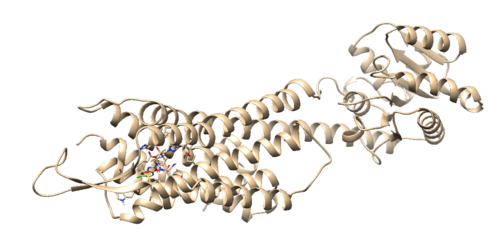
On the top in the box that reads Prmtop, press the browse button and choose the 4s0v.wet.complex.parm7. Next, click in the grey box and click on the 'Add' button. Then choose the 4s0v.wet.complex.rst7 file. Finally click the 'OK' button. Once the file is loaded you should see your ligand in the binding site of your protein:
Everything looks good so we can move onto the next step.
Complex Equilibration
Before we can run our structure through the AMBER program we need to minimize it and make sure it's at equilibrium. We will be working on Seawulf, please cd into your 004.equilibration directory. Relaxing the structure involves creating nine input files with the following names and commands:
Create 01.min.mdin
vi 01.min.mdin
Copy the following lines:
Minmize all the hydrogens &cntrl imin=1, ! Minimize the initial structure maxcyc=5000, ! Maximum number of cycles for minimization ntb=1, ! Constant volume ntp=0, ! No pressure scaling ntf=1, ! Complete force evaluation ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask=":1-131 & !@H=", ! atoms to be restrained restraint_wt=5.0, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format /
Create 02.equil.mdin
vi 02.equil.mdin
Copy the following lines:
MD simualation &cntrl imin=0, ! Perform MD nstlim=50000 ! Number of MD steps ntb=2, ! Constant Pressure ntc=1, ! No SHAKE on bonds between hydrogens dt=0.001, ! Timestep (ps) ntp=1, ! Isotropic pressure scaling barostat=1 ! Berendsen taup=0.5 ! Pressure relaxtion time (ps) ntf=1, ! Complete force evaluation ntt=3, ! Langevin thermostat gamma_ln=2.0 ! Collision Frequency for thermostat ig=-1, ! Random seed for thermostat temp0=298.15 ! Simulation temperature (K) ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask=":1-131 & !@H=", ! atoms to be restrained restraint_wt=5.0, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format iwrap=1, ! iwrap is turned on /
Create 03.min.mdin
vi 03.min.mdin
Copy the following lines:
Minmize all the hydrogens &cntrl imin=1, ! Minimize the initial structure maxcyc=1000, ! Maximum number of cycles for minimization ntb=1, ! Constant volume ntp=0, ! No pressure scaling ntf=1, ! Complete force evaluation ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask=":1-131 & !@H=", ! atoms to be restrained restraint_wt=2.0, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format /
Create 04.min.mdin
vi 04.min.mdin
Copy the following lines:
Minmize all the hydrogens &cntrl imin=1, ! Minimize the initial structure maxcyc=1000, ! Maximum number of cycles for minimization ntb=1, ! Constant volume ntp=0, ! No pressure scaling ntf=1, ! Complete force evaluation ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask=":1-131 & !@H=", ! atoms to be restrained restraint_wt=0.1, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format /
Create 05.min.mdin
vi 05.min.mdin
Copy the following lines:
Minmize all the hydrogens &cntrl imin=1, ! Minimize the initial structure maxcyc=1000, ! Maximum number of cycles for minimization ntb=1, ! Constant volume ntp=0, ! No pressure scaling ntf=1, ! Complete force evaluation ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask=":1-131 & !@H=", ! atoms to be restrained restraint_wt=0.05, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format /
Create 06.equil.mdin
vi 06.equil.mdin
Copy the following lines:
MD simualation &cntrl imin=0, ! Perform MD nstlim=50000 ! Number of MD steps ntb=2, ! Constant Pressure ntc=1, ! No SHAKE on bonds between hydrogens dt=0.001, ! Timestep (ps) ntp=1, ! Isotropic pressure scaling barostat=1 ! Berendsen taup=0.5 ! Pressure relaxtion time (ps) ntf=1, ! Complete force evaluation ntt=3, ! Langevin thermostat gamma_ln=2.0 ! Collision Frequency for thermostat ig=-1, ! Random seed for thermostat temp0=298.15 ! Simulation temperature (K) ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask=":1-131 & !@H=", ! atoms to be restrained restraint_wt=1.0, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format iwrap=1, ! iwrap is turned on /
Create 07.equil.mdin
vi 07.equil.mdin
Copy the following lines:
MD simulation &cntrl imin=0, ! Perform MD nstlim=50000 ! Number of MD steps ntx=5, ! Positions and velocities read formatted irest=1, ! Restart calculation ntc=1, ! No SHAKE on for bonds with hydrogen dt=0.001, ! Timestep (ps) ntb=2, ! Constant Pressure ntp=1, ! Isotropic pressure scaling barostat=1 ! Berendsen taup=0.5 ! Pressure relaxtion time (ps) ntf=1, ! Complete force evaluation ntt=3, ! Langevin thermostat gamma_ln=2.0 ! Collision Frequency for thermostat ig=-1, ! Random seed for thermostat temp0=298.15 ! Simulation temperature (K) ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask=":1-131 & !@H=", ! atoms to be restrained restraint_wt=0.5, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format iwrap=1, ! iwrap is turned on /
Create 08.equil.mdin
vi 08.equil.mdin
Copy the following lines:
MD simulations &cntrl imin=0, ! Perform MD nstlim=50000 ! Number of MD steps ntx=5, ! Positions and velocities read formatted irest=1, ! Restart calculation ntc=1, ! No SHAKE on for bonds with hydrogen dt=0.001, ! Timestep (ps) ntb=2, ! Constant Pressure ntp=1, ! Isotropic pressure scaling barostat=1 ! Berendsen taup=0.5 ! Pressure relaxtion time (ps) ntf=1, ! Complete force evaluation ntt=3, ! Langevin thermostat gamma_ln=2.0 ! Collision Frequency for thermostat ig=-1, ! Random seed for thermostat temp0=298.15 ! Simulation temperature (K) ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask=":1-131@CA,C,N", ! atoms to be restrained restraint_wt=0.1, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format iwrap=1, ! iwrap is turned on /
Create 09.equil.mdin
vi 09.equil.mdin
Copy the following lines:
MD simulations &cntrl imin=0, ! Perform MD nstlim=50000 ! Number of MD steps ntx=5, ! Positions and velocities read formatted irest=1, ! Restart calculation ntc=1, ! No SHAKE on for bonds with hydrogen dt=0.001, ! Timestep (ps) ntb=2, ! Constant Pressure ntp=1, ! Isotropic pressure scaling barostat=1 ! Berendsen taup=0.5 ! Pressure relaxtion time (ps) ntf=1, ! Complete force evaluation ntt=3, ! Langevin thermostat gamma_ln=2.0 ! Collision Frequency for thermostat ig=-1, ! Random seed for thermostat temp0=298.15 ! Simulation temperature (K) ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask=":1-131@CA,C,N", ! atoms to be restrained restraint_wt=0.1, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format iwrap=1, ! iwrap is turned on /
Once you have all nine files in your directory we need to create a slurm script, 4s0v.equilibration.slurm, to run them:
#!/bin/bash # #SBATCH --job-name=4s0v_equilibration #SBATCH --output=equilibration_output.txt #SBATCH --ntasks-per-node=96 #SBATCH --nodes=6 #SBATCH --time=48:00:00 #SBATCH -p long-96core do_parallel="mpirun pmemd.MPI"
prmtop="../003.tleap/4s0v.wet.complex.parm7" coords="../003.tleap/4s0v.wet.complex.rst7" MDINPUTS=(01.min 02.equil 03.min 04.min 05.min 06.equil 07.equil 08.equil 09.equil)
for input in ${MDINPUTS[@]}; do
$do_parallel -O -i ${input}.mdin -o ${input}.mdout -p $prmtop -c ${coords}.rst7 -ref ${coords}.rst7 -x ${input}.trj -inf ${input}.info -r ${input}.rst7
coords=$input
done
Submit the above file to Milan (or Seawulf) by typing:
sbatch 4s0v.sequilibration.slurm