Difference between revisions of "2023 DOCK tutorial 2 with PDBID 3WZE"
Stonybrook (talk | contribs) (→Energy minimization) |
Stonybrook (talk | contribs) (→Reproducing the PDB's binding with DOCK) |
||
| Line 266: | Line 266: | ||
= '''Reproducing the PDB's binding with DOCK''' = | = '''Reproducing the PDB's binding with DOCK''' = | ||
| + | Change the current directory to perform DOCK: | ||
| + | cd 004_dock | ||
==='''Rigid Docking'''=== | ==='''Rigid Docking'''=== | ||
This is computationally least expensive since the ligand will be stable, no rotational degrees will be taken into consideration. The energy minimized ligand mol2 file will be used in this simulation. First we will need to create a new file “rigid.in”: | This is computationally least expensive since the ligand will be stable, no rotational degrees will be taken into consideration. The energy minimized ligand mol2 file will be used in this simulation. First we will need to create a new file “rigid.in”: | ||
| Line 282: | Line 284: | ||
orient_ligand yes | orient_ligand yes | ||
automated_matching yes | automated_matching yes | ||
| − | receptor_site_file selected_spheres.sph | + | receptor_site_file ../002_surface_spheres/selected_spheres.sph |
max_orientations 1000 | max_orientations 1000 | ||
critical_points no | critical_points no | ||
| Line 296: | Line 298: | ||
grid_score_vdw_scale 1 | grid_score_vdw_scale 1 | ||
grid_score_es_scale 1 | grid_score_es_scale 1 | ||
| − | grid_score_grid_prefix grid | + | grid_score_grid_prefix ../003_gridbox/grid |
multigrid_score_secondary no | multigrid_score_secondary no | ||
dock3.5_score_secondary no | dock3.5_score_secondary no | ||
| Line 350: | Line 352: | ||
internal_energy_rep_exp 12 | internal_energy_rep_exp 12 | ||
internal_energy_cutoff 100.0 | internal_energy_cutoff 100.0 | ||
| − | ligand_atom_file 3WZE_lig.mol2 | + | ligand_atom_file ../001_structure/3WZE_lig.mol2 |
limit_max_ligands no | limit_max_ligands no | ||
skip_molecule no | skip_molecule no | ||
| Line 356: | Line 358: | ||
calculate_rmsd yes | calculate_rmsd yes | ||
use_rmsd_reference_mol yes | use_rmsd_reference_mol yes | ||
| − | rmsd_reference_filename 3WZE_lig.mol2 | + | rmsd_reference_filename ../001_structure/3WZE_lig.mol2 |
use_database_filter no | use_database_filter no | ||
orient_ligand no | orient_ligand no | ||
| Line 439: | Line 441: | ||
orient_ligand yes | orient_ligand yes | ||
automated_matching yes | automated_matching yes | ||
| − | receptor_site_file selected_spheres.sph | + | receptor_site_file ../002_surface_spheres/selected_spheres.sph |
max_orientations 1000 | max_orientations 1000 | ||
critical_points no | critical_points no | ||
Revision as of 13:41, 14 March 2023
In this tutorial, you will learn how use the program DOCK6.10 to perform a virtual screen, in which you assess how well the molecules in a library of drug-like molecules bind to a protein of known structure.
Contents
Introduction
A protein whose function is found to be involved in one or more diseases may become a target for pharmaceutical design. Oftentimes, these pharmaceuticals are designed to compete with the enzyme's native substrate for the enzyme's active site, making many pharmaceutical molecules competitive inhibitors of their protein targets. If the target protein's structure is known, and the active site can be identified, then performing a virtual screen can be a monetarily and temporally efficient method of identifying molecules which are likely to bind well to the target's active site.
A virtual screen is set up by first preparing the enzyme's structure and the structure of its native substrate for docking, then the residues important for the native ligand to bind are identified by generating a footprint. A large library of drug-like molecules is then downloaded from a database such as ZINC (https://zinc.docking.org/), and, using the footprint and enzyme structure, docked into the enzyme using a program such as DOCK6.10. Results are then assessed to see which drug-like compounds match the native substrate's footprint profile and which are energetically comfortable within the simulated active site. Such molecules could then be tested biochemically for their ability to inhibit the target protein, sparing biochemists the hassle of having to test hundreds of thousands of compounds in physical screening experiments.
Software and Files
PDB 3WZE
PDB stands for "Protein Data Bank", which is a repository for the experimentally solved structures of proteins. Each protein structure is assigned a 4 digit code, and 3WZE is the code assigned to the solved structure of vascular endothelial growth factor receptor 2 (VEGFR2) bound to the inhibitor sorafenib. VEGFR2 is a receptor tyrosine kinase, meaning that it is an integral membrane protein that has an exofacial receptor domain, transmembrane domain, and cytofacial kinase domain. Because of the difficulties of protein purification, crystallization, and structure solution, many protein structures in the protein data bank are incomplete: lacking regions that are intrinsically disordered or otherwise not conducive to crystallization. PDB 3WZE is one such structure, because it only contains structural data for the cytofacial kinase domain of VEGFR2.
The active site of kinases is well characterized, and sorafenib is shown bound within it in PDB 3WZE, which will be useful for conducting later steps in the virtual screen.
Download the .pdb file of 3WZE, and use a program such as Chimera or ChimeraX to open and view it.
The Protein Data Bank home page can be found here:
DOCK6.10
DOCK is a program used to model conformations of molecules, and is typically used to find low potential energy conformations of drug-like molecules within the active sites of protein structures. Because molecules in nature tend to adopt conformations of the lowest possible potential energy, identifying low-energy conformations of molecules within proteins can be a useful way of identifying how such molecules may bind in nature. Thus, instead of having to manually test out libraries of thousands of molecules to identify potential therapeutics, a library of drug-like molecules can be docked into a protein of interest, vastly expediting the medicine development process at one of its earliest steps.
DOCK can be used for more purposes than drug identification though, and is capable of rigid and flexible docking, de novo design, and genetic algorithms. Rigid docking is when DOCK identifies low-potential energy conformations of a ligand within the target protein, but while keeping the ligand rigid. Flexible docking on the other hand allows for rotations around the ligand's internal rotatable bonds. De novo design is when DOCK attempts to build a new ligand outwards from a starting anchor specified by the user. Finally, genetic algorithms generate new ligands similarly tpo de novo design, but it also utilizes the principles of Darwinian evolution and natural selection to select for effective ligands over the course of multiple generations of ligand development.
For the purposes of this tutorial, you will only be using the rigid and flexible docking capabilities of DOCK. For more information on the other uses of DOCK, check the other tutorials in this wiki or consult the DOCK 6.10 User Manual:
https://dock.compbio.ucsf.edu/DOCK_6/dock6_manual.htm#RigidandFlexibleLigandDocking
Chimera
Chimera is a visualization program that can display the 3-dimensional structures of molecules. Although Chimera is no longer actively developed, it remains a useful tool and an essential part of this tutorial. As you follow this tutorial, molecular visualization programs like Chimera and ChimeraX will be useful for preparing the system for the virtual screen and checking the results of each step. The home page of Chimera can be found here:
https://www.cgl.ucsf.edu/chimera/
ChimeraX (optional)
Chimera is now no longer actively developed, and has been succeeded by ChimeraX, which is developed by the same group. Although ChimeraX has lost some of the functionality of its predecessor, it has new capabilities to compensate, and it is easier to operate using typed commands, whereas Chimera requires clicking through menus. That being said, Chimera is still required for this tutorial because ChimeraX cannot open .sph files and it cannot save a surface as a .dms file.
Separate instructions for completing the tutorial with ChimeraX will be provided alongside the instructions for using Chimera in each section.
The home page for ChimeraX can be found here:
https://www.cgl.ucsf.edu/chimerax/
Alphafold (optional)
Alphafold is a protein structure prediction program from Google's Deepmind, and it was the first program to predict the structures of proteins in the annual CASP competition to within 90% accuracy in 2020 (https://predictioncenter.org/casp14/zscores_final.cgi). Using this program, one can generate a reasonably accurate prediction of a protein's structure using only its amino acid sequence. In the context of virtual screening, this means that a protein's structure no longer needs to be solved experimentally before one can embark on a virtual screen of the target. As long as the active site can be identified (which is often done by comparing the predicted structure to homologous proteins with solved structures), one can perform a virtual screen of a protein of unsolved structure.
Even without a university server, Alphafold can be used from within ChimeraX by going to Tools -> Structure Prediction -> Alphafold. This will bring up a menu in which you can paste an amino acid sequence for prediction, searching, or retrieval from the Alphafold database. All human proteins have already been predicted by Alphafold, and their structures can be easily retrieved using the protein's UniProt identifier and the Fetch button. Non-human proteins will have to be predicted from scratch by inputting their amino acid sequence and using the Predict button.
Because this tutorial will use PDB file 3WZE, which contains the solved structure of the Vascular Endothelial Growth Factor Receptor and a bound inhibitor called sorafenib, Alphafold will be unnecessary for this tutorial, but the broadened scope of what virtual screens are possible as a result of this program is worth noting nonetheless.
The Alphafold home page can be found here:
Using the Terminal
Seawulf uses the Linux language for various commands. Here are some of the basic commands that will make running the tutorial run smoothly.
The first command is cd, which is used to change into different directories. To go to a desired directory do:
cd /desired/directory
To move to a previous directory you were just in, type:
cd ../
The cd command can also be used to get into a folder, type:
cd /desired/folder
The next command is the ls command. This will list out all the files or folders in the directory you are currently in, type:
ls
To make a new directory, type:
mdkir name_of_directory
To remove a file, type:
rm file
To delete a directory and all its contents, type:
rm -r directory
To create a file in the terminal, type:
vi filename
This will open up a blank screen in which you can start typing. To edit the file, first type “i”. This will make the environment go to “INSERT” mode. You can now freely type. In order to save whatever you wrote in the file, click the escape button. This will remove you from “INSERT” mode. Next type:
:wq
This will save any changes you have made to your file
To move a file to a new location, type:
mv file /new/destination
To make a copy of a file, type:
cp file file_new_name
This command can also be used to copy directories as well, type:
cp -r directory_1 directory_2
Directory Organization
Having defined folders established before starting is a great way to maintain organization and clarity as you go.
000_files 001_structure 002_surface_spheres 003_gridbox 004_dock 005_virtual_screen 006_virtual_screen_mpi 007_cartesian_min 008_rescore
Additionally, file names should be clear and logical. We recommend starting with the PDB code followed by the receptor or ligand and ending with what changes were made. For example, 3WZE_lig_AddH_addCharge.mol2 indicates that this file is the ligand for 3WZE and hydrogens and charge have been added. Also, bear in mind that one should use underscores instead of spaces when naming files and directories. Otherwise, spaces in file or directory names can confuse many linux commands and make them mistake the names for command arguments.
Preparing the Receptor
Preparing the Ligand
Surface & Spheres
Surface Generation
The generation of the surface of the receptor protein will have better visualization of the binding site of the receptor. In order to do so, load the mol2 file of 3WZE receptor in Chimera:
Action < Surface < show
Then we will save the dms(molecular surface) file as 3WZE_rec.dms, which will be used in the next step of spheres generation:
Tools < Structure Editing < Write DMS
Change current working directory in terminal for sphere generation and selection:
cd 002_surface_spheres
Spheres Generation
First create a new file “INSPH” by typing the following into terminal:
vi INSPH
Then type the following information into this file (anything after # are comments that are references for reading only, and should not be putting into the file) :
3WZE_rec.dms #Molecular Surface file R #This line specifies whether to make the spheres in the model’s exterior (R) or interior (L) X #Specifies that the surface points from the dms file should be used for making spheres(?) 0 #Minimum radius between spheres 4.0 #Max radius of each sphere 1.4 #Minimum radius of each sphere 3WZE.sph #Name of the output file that will be made
Then save the file and run the following command on terminal: note that if you want to re-run this command, make sure you remove file “OUTSPH” before your re-run because this file cannot be overwritten.
sphgen -i INSPH -o OUTSPH
The output file “3WZE.sph” can be visualized by using Chimera, load the output file along with the receptor protein mol2 file, the two structures should be overlapping:
Spheres selection
Since we are only interested in what’s happening in the binding site, we will remove spheres that are far away from the ligand. To do so, use the following command and an output file “selected_spheres.sph” will be generated:
sphere_selector 3WZE.sph 3WZE_lig.mol2 10.0 #sphere_selector [sphere output file] [ligand mol2 file] [radius for distance from ligand]
Box and Grid
Change the current working directory in terminal for box and grid generation:
cd 003_gridbox
Box generation
First we will generate a box that will be used to generate the grid. To generate a box, we will need to create a new file “showbox.in” by using the following command:
vi showbox.in
Then type the following information into the file (anything after # are comments that are references for reading only, and should not be putting into the file) :
Y #Indicates that we want to make a box 8.0 #Extra length beyond the spheres to be included by the box in all directions selected_spheres.sph #Indicates that the box should be constructed to encompass the spheres in this file. 1 #This specifies which cluster of spheres should be used to generate the box. 3WZE.box.pdb #Name of output file
Then, run the following code to get the boxed structure:
showbox < showbox.in
Grid generation
Create a new file “grid.in” by using the following command:
vi grid.in
Then type the following information into the file:
compute_grids yes grid_spacing 0.4 output_molecule no contact_score no energy_score yes energy_cutoff_distance 9999 atom_model a attractive_exponent 6 repulsive_exponent 9 distance_dielectric yes dielectric_factor 4 bump_filter yes bump_overlap 0.75 receptor_file 3WZE_rec.mol2 box_file 3WZE.box.pdb vdw_definition_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_AMBER_parm99.defn score_grid_prefix grid
Then generate the grid by typing the following command:
grid -i grid.in -o gridinfo.out
This will generate three files: “gridinfo.out”, “grid.bmp”, and “grid.nrg”. Open “gridinfo.out” and check if the information about the receptor matches the that from the paper and your previous preparations. Also check to make sure there are no error messages at the bottom.
Energy minimization
Change the current working directory in terminal to do preparation for DOCK:
cd 004_dock
To improve the accuracy of DOCK calculation of ligand protein binding, we will have to minimize the potential energy of the ligand. To do so, create a new file “min.in” by using the following command:
vi min.in
Then type the following information into the file:
conformer_search_type rigid use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 ligand_atom_file ../000_files/3WZE_rec.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd yes use_rmsd_reference_mol yes rmsd_reference_filename ../000_files/3WZE_rec.mol2 use_database_filter no orient_ligand no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary yes grid_score_secondary no grid_score_rep_rad_scale 1 grid_score_vdw_scale 1 grid_score_es_scale 1 grid_score_grid_prefix grid multigrid_score_secondary no dock3.5_score_secondary no continuous_score_secondary no footprint_similarity_score_secondary no pharmacophore_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand yes simplex_max_iterations 1000 simplex_tors_premin_iterations 0 simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1.0 simplex_trans_step 1.0 simplex_rot_step 0.1 simplex_tors_step 10.0 simplex_random_seed 0 simplex_restraint_min yes simplex_coefficient_restraint 10.0 atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl ligand_outfile_prefix 3WZE.lig.min write_orientations no num_scored_conformers 1 rank_ligands no
Then run the following command through the use of DOCK:
dock6 -i min.in
The output file “3WZE.lig.min_scored.mol2” is the potential energy minimized version of the ligand.
Reproducing the PDB's binding with DOCK
Change the current directory to perform DOCK:
cd 004_dock
Rigid Docking
This is computationally least expensive since the ligand will be stable, no rotational degrees will be taken into consideration. The energy minimized ligand mol2 file will be used in this simulation. First we will need to create a new file “rigid.in”:
vi rigid.in
Type the following information into the new file:
conformer_search_type rigid use_internal_energy yes ligand_atom_file 3WZE.lig.min_scored.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd yes use_rmsd_reference_mol yes rmsd_reference_filename 3WZE.lig.min_scored.mol2 use_database_filter no orient_ligand yes automated_matching yes receptor_site_file ../002_surface_spheres/selected_spheres.sph max_orientations 1000 critical_points no chemical_matching no use_ligand_spheres no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary yes grid_score_secondary no grid_score_rep_rad_scale 1 grid_score_vdw_scale 1 grid_score_es_scale 1 grid_score_grid_prefix ../003_gridbox/grid multigrid_score_secondary no dock3.5_score_secondary no continuous_score_secondary no footprint_similarity_score_secondary no pharmacophore_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand yes simplex_max_iterations 1000 simplex_tors_premin_iterations 0 simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1.0 simplex_trans_step 1.0 simplex_rot_step 0.1 simplex_tors_step 10.0 simplex_random_seed 0 simplex_restraint_min no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl ligand_outfile_prefix rigid.out write_orientations no num_scored_conformers 1 rank_ligands no
Then run the following command:
dock6 -i rigid.in -o rigid.out
If the inputs are all correct, there will be a new output file “rigid.out_scored.mol2”.
Fixed Anchor Docking
In fixed anchor docking, most part of the ligand will remain fixed, and the rest will have some rotational freedom. First create a new file “fixed.in”:
vi fixed.in
Then type the following into the file:
conformer_search_type flex user_specified_anchor no limit_max_anchors no min_anchor_size 5 pruning_use_clustering yes pruning_max_orients 1000 pruning_clustering_cutoff 100 pruning_conformer_score_cutoff 100.0 pruning_conformer_score_scaling_factor 1.0 use_clash_overlap no write_growth_tree no write_fragment_libraries no use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 ligand_atom_file ../001_structure/3WZE_lig.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd yes use_rmsd_reference_mol yes rmsd_reference_filename ../001_structure/3WZE_lig.mol2 use_database_filter no orient_ligand no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary yes grid_score_secondary no grid_score_rep_rad_scale 1 grid_score_vdw_scale 1 grid_score_es_scale 1 grid_score_grid_prefix grid multigrid_score_secondary no dock3.5_score_secondary no continuous_score_secondary no footprint_similarity_score_secondary no pharmacophore_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand yes minimize_anchor yes minimize_flexible_growth yes use_advanced_simplex_parameters no simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1.0 simplex_trans_step 1.0 simplex_rot_step 0.1 simplex_tors_step 10.0 simplex_anchor_max_iterations 500 simplex_grow_max_iterations 500 simplex_grow_tors_premin_iterations 0 simplex_random_seed 0 simplex_restraint_min no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl ligand_outfile_prefix fixed.out write_orientations no num_scored_conformers 100 write_conformations no cluster_conformations yes cluster_rmsd_threshold 2.0 rank_ligands no
Then run the following command:
dock6 -i fixed.in -o fixed.out
If the inputs are all correct, there will be a new output file “fixed.out_scored.mol2”.
Flexible Docking
Flexible docking is the computationally most expensive out of the three types of docking we introduced. Full rotational freedom and internal degrees of freedom are being considered. First creat a new file “flex.in”:
vi flex.in
Type the following information into the file:
conformer_search_type flex user_specified_anchor no limit_max_anchors no min_anchor_size 5 pruning_use_clustering yes pruning_max_orients 1000 pruning_clustering_cutoff 100 pruning_conformer_score_cutoff 100.0 pruning_conformer_score_scaling_factor 1.0 use_clash_overlap no write_growth_tree no write_fragment_libraries no use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 ligand_atom_file 3WZE.lig.min_scored.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd yes use_rmsd_reference_mol yes rmsd_reference_filename 3WZE.lig.min_scored.mol2 use_database_filter no orient_ligand yes automated_matching yes receptor_site_file ../002_surface_spheres/selected_spheres.sph max_orientations 1000 critical_points no chemical_matching no use_ligand_spheres no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary yes grid_score_secondary no grid_score_rep_rad_scale 1 grid_score_vdw_scale 1 grid_score_es_scale 1 grid_score_grid_prefix grid multigrid_score_secondary no dock3.5_score_secondary no continuous_score_secondary no footprint_similarity_score_secondary no pharmacophore_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand yes minimize_anchor yes minimize_flexible_growth yes use_advanced_simplex_parameters no simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1.0 simplex_trans_step 1.0 simplex_rot_step 0.1 simplex_tors_step 10.0 simplex_anchor_max_iterations 500 simplex_grow_max_iterations 500 simplex_grow_tors_premin_iterations 0 simplex_random_seed 0 simplex_restraint_min no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl ligand_outfile_prefix flex.out write_orientations no num_scored_conformers 1 rank_ligands no
Then run the following command:
dock6 -i flex.in -o flex.out
If the inputs are all correct, there will be a new output file “flex.out_scored.mol2”.
Footprinting
Dock6 has a feature in which we can visualize both the electrostatic and Van der interactions between the ligand and protein. This will show which amino acids are contributing to the binding of the ligand to the protein.
To determine the molecular footprint, create the following input file in your 004_dock directory:
vi footprint.in
Copy the following into your file:
conformer_search_type rigid use_internal_energy no ligand_atom_file 3WZE.lig.min_scored.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd no use_database_filter no orient_ligand no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary no grid_score_secondary no multigrid_score_primary no multigrid_score_secondary no dock3.5_score_primary no dock3.5_score_secondary no continuous_score_primary no continuous_score_secondary no footprint_similarity_score_primary yes footprint_similarity_score_secondary no fps_score_use_footprint_reference_mol2 yes fps_score_footprint_reference_mol2_filename ../001_structure/3wze.lig.wH.am1bcc.mol2 fps_score_foot_compare_type Euclidean fps_score_normalize_foot no fps_score_foot_comp_all_residue yes fps_score_receptor_filename ../001_structure/3wze_rec_wH.mol2 fps_score_vdw_att_exp 6 fps_score_vdw_rep_exp 9 fps_score_vdw_rep_rad_scale 1 fps_score_use_distance_dependent_dielectric yes fps_score_dielectric 4.0 fps_score_vdw_fp_scale 1 fps_score_es_fp_scale 1 fps_score_hb_fp_scale 0 pharmacophore_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl ligand_outfile_prefix footprint.out write_footprints yes write_hbonds yes write_orientations no num_scored_conformers 1 rank_ligands no
In order to run the footprint calculation, type:
dock6 -i footprint.in -o footprint.out
An error might occur when running footprint with dock, but that is fine. Three files will be generated after a successful run. The files that are generated are footprint.out_footprint_scored.txt footprint.out_hbond_scored.txt footprint.out_scored.mol2. In order to visualize the molecular footprint we need to use a python script provided here a prewritten python script. To run the script just do:
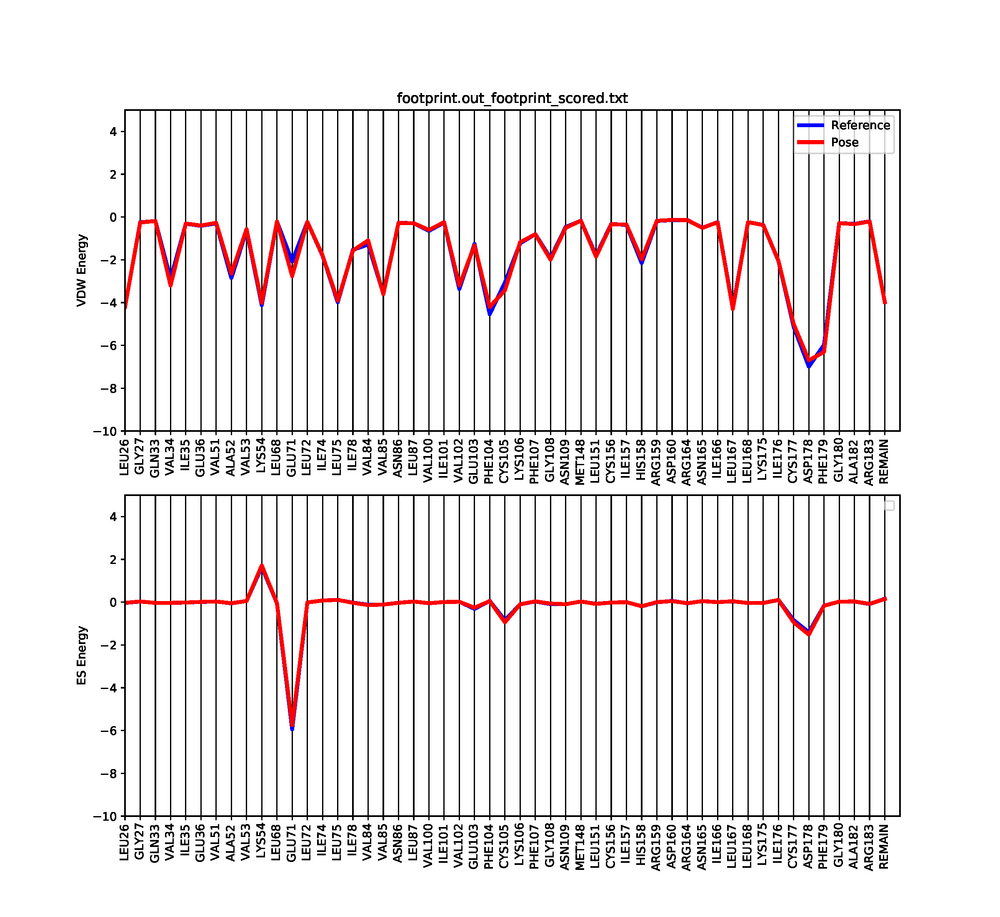
python plot_footprint_single_magnitude.py footprint.out_footprint_scored.txt 50
This python script will generate a graph that shows the Van der Waals and Electrostatic contributions of the 50 most contributing amino acids. Use a pdf viewer to visualize the graph you just created. It should look similar to this:
Virtual Screen
By having a predetermined compound library, virtual screening in DOCK can score these in the compound library based on the input parameters. In this case, we used a smaller library “VS_library_25K.mol2” that was provided by the instructor . First create a new file “virtual.in” for inputting virtual screening parameters:
vi virtual.in
Then type the following information into the file:
conformer_search_type flex user_specified_anchor no limit_max_anchors no min_anchor_size 5 pruning_use_clustering yes pruning_max_orients 1000 pruning_clustering_cutoff 100 pruning_conformer_score_cutoff 100.0 pruning_conformer_score_scaling_factor 1.0 use_clash_overlap no write_growth_tree no write_fragment_libraries no use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 ligand_atom_file VS_library_25K.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd no use_database_filter no orient_ligand yes automated_matching yes receptor_site_file selected_spheres.sph max_orientations 1000 critical_points no chemical_matching no use_ligand_spheres no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary yes grid_score_secondary no grid_score_rep_rad_scale 1 grid_score_vdw_scale 1 grid_score_es_scale 1 grid_score_grid_prefix grid multigrid_score_secondary no dock3.5_score_secondary no continuous_score_secondary no footprint_similarity_score_secondary no pharmacophore_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand yes minimize_anchor yes minimize_flexible_growth yes use_advanced_simplex_parameters no simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1.0 simplex_trans_step 1.0 simplex_rot_step 0.1 simplex_tors_step 10.0 simplex_anchor_max_iterations 500 simplex_grow_max_iterations 500 simplex_grow_tors_premin_iterations 0 simplex_random_seed 0 simplex_restraint_min no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl ligand_outfile_prefix virtual.out write_orientations no num_scored_conformers 1 rank_ligands no
To prevent overload of the current working load and speed up the process, we can write the code for virtual screening along with requesting more nodes in s shell script and submit the job. To do so, first create a new file “virtual.sh”:
vi virtual.sh
Then type the following information into this file:
#!/bin/bash #SBATCH --time=5:00:00 #SBATCH --nodes=4 #SBATCH --ntasks=112 #SBATCH --job-name=3WZE_flex_dock #SBATCH --output=3WZE_flex_dock #SBATCH -p long-28core mpirun -np 112 dock6.mpi -i virtual.in -o 3WZE.virtual.mpi.out
Then submit the job by typing the following command:
sbatch virtual.sh
Since we specified the number of task to be 112, we are expecting to see 112 output files, name in the format "virtual.out.xxx". In addition, the output contains a mol2 file with all 112 virtual screening results inside.
Cartesian Energy Minimization
We have to perform energy minimization on the virtual screening results before analyzing. First, create a new file “cart_min.in”:
vi cart_min.in
Then type the following information into the file:
conformer_search_type rigid use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 ligand_atom_file 3WZE_virtual.out_scored.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd no use_database_filter no orient_ligand no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary no grid_score_secondary no multigrid_score_primary no multigrid_score_secondary no dock3.5_score_primary no dock3.5_score_secondary no continuous_score_primary yes continuous_score_secondary no cont_score_rec_filename 3WZE_rec.mol2 cont_score_att_exp 6 cont_score_rep_exp 12 cont_score_rep_rad_scale 1 cont_score_use_dist_dep_dielectric yes cont_score_dielectric 4.0 cont_score_vdw_scale 1 cont_score_es_scale 1 footprint_similarity_score_secondary no pharmacophore_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand yes simplex_max_iterations 1000 simplex_tors_premin_iterations 0 simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1.0 simplex_trans_step 1.0 simplex_rot_step 0.1 simplex_tors_step 10.0 simplex_random_seed 0 simplex_restraint_min no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl ligand_outfile_prefix 3WZE.virtualscreen_cart.min write_orientations no num_scored_conformers 1 rank_ligands no
Then we write a new shell script “cart_min.sh” to request node and run the code to prevent overflow:
vi cart_min.sh
The following lines should be included in the shell file:
#!/bin/bash #SBATCH --time=10:00:00 #SBATCH --nodes=4 #SBATCH --ntasks=112 #SBATCH --job-name=3WZE_flex_dock #SBATCH --output=3WZE_flex_dock #SBATCH -p long-28core mpirun -np 112 dock6.mpi -i cart_min.in -o 3WZE_cart_min.virtual.mpi.out
Then submit the job by running the following command:
sbatch cart_min.sh
We then will be able to see the results.
Rescoring the Virtual Screen
We can rescore the energy minimized virtual screening results. This can rank the results based on user defined molecular properties. To do so, make a new file “rescore.in”:
vi rescore.in
Then type the following information into the file:
conformer_search_type rigid use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 ligand_atom_file 3WZE.vs.min_scored.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd no use_database_filter no orient_ligand no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary no grid_score_secondary no multigrid_score_primary no multigrid_score_secondary no dock3.5_score_primary no dock3.5_score_secondary no continuous_score_primary no continuous_score_secondary no footprint_similarity_score_primary no footprint_similarity_score_secondary no pharmacophore_score_primary no pharmacophore_score_secondary no descriptor_score_primary yes descriptor_score_secondary no descriptor_use_grid_score no descriptor_use_multigrid_score no descriptor_use_continuous_score no descriptor_use_footprint_similarity yes descriptor_use_pharmacophore_score yes descriptor_use_tanimoto yes descriptor_use_hungarian yes descriptor_use_volume_overlap yes descriptor_fps_score_use_footprint_reference_mol2 yes descriptor_fps_score_footprint_reference_mol2_filename 3wze.lig.min_scored.mol2 descriptor_fps_score_foot_compare_type Euclidean descriptor_fps_score_normalize_foot no descriptor_fps_score_foot_comp_all_residue yes descriptor_fps_score_receptor_filename 3wze_rec.mol2 descriptor_fps_score_vdw_att_exp 6 descriptor_fps_score_vdw_rep_exp 12 descriptor_fps_score_vdw_rep_rad_scale 1 descriptor_fps_score_use_distance_dependent_dielectric yes descriptor_fps_score_dielectric 4.0 descriptor_fps_score_vdw_fp_scale 1 descriptor_fps_score_es_fp_scale 1 descriptor_fps_score_hb_fp_scale 0 descriptor_fms_score_use_ref_mol2 yes descriptor_fms_score_ref_mol2_filename 3wze.lig.min_scored.mol2 descriptor_fms_score_write_reference_pharmacophore_mol2 no descriptor_fms_score_write_reference_pharmacophore_txt no descriptor_fms_score_write_candidate_pharmacophore no descriptor_fms_score_write_matched_pharmacophore no descriptor_fms_score_compare_type overlap descriptor_fms_score_full_match yes descriptor_fms_score_match_rate_weight 5.0 descriptor_fms_score_match_dist_cutoff 1.0 descriptor_fms_score_match_proj_cutoff 0.7071 descriptor_fms_score_max_score 20 descriptor_fingerprint_ref_filename 3wze.lig.min_scored.mol2 descriptor_hms_score_ref_filename 3wze.lig.min_scored.mol2 descriptor_hms_score_matching_coeff -5 descriptor_hms_score_rmsd_coeff 1 descriptor_volume_score_reference_mol2_filename 3wze.lig.min_scored.mol2 descriptor_volume_score_overlap_compute_method analytical descriptor_weight_fps_score 1 descriptor_weight_pharmacophore_score 1 descriptor_weight_fingerprint_tanimoto -1 descriptor_weight_hms_score 1 descriptor_weight_volume_overlap_score -1 gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl chem_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/chem.defn pharmacophore_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/ph4.defn ligand_outfile_prefix descriptor.out write_footprints yes write_hbonds yes write_orientations no num_scored_conformers 1 rank_ligands no
To prevent overload, we write a new shell script to request nodes and run the line:
vi rescore.sh
Type the following information into the file:
#!/bin/bash #SBATCH --time=10:00:00 #SBATCH --nodes=4 #SBATCH --ntasks=112 #SBATCH --job-name=3WZE_flex_dock #SBATCH --output=3WZE_flex_dock #SBATCH -p long-28core mpirun -np 112 dock6.mpi -i rescore.in -o rescore.out
Then run the following command:
sbatch rescore.sh
There will be a file “descriptor.out_scored.mol2” when the job is done. We can download the file to local and perform visual analysis by using Chimera.