Difference between revisions of "2015 DOCK tutorial with Poly(ADP-ribose) polymerase (PARP)"
Stonybrook (talk | contribs) (→Placing Spheres) |
Stonybrook (talk | contribs) (→Placing Spheres) |
||
| Line 94: | Line 94: | ||
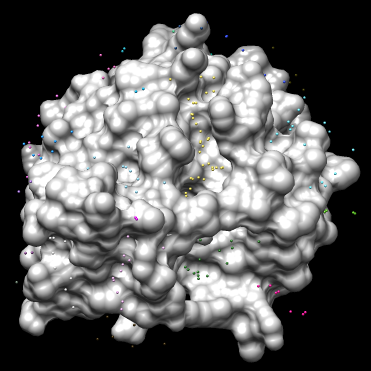
You should have an image like this: | You should have an image like this: | ||
[[Image:4tkg.rec_allsph.png|thumb|center|375px|4TKG Receptor surface (light gray) with all spheres (various colors) generated]] | [[Image:4tkg.rec_allsph.png|thumb|center|375px|4TKG Receptor surface (light gray) with all spheres (various colors) generated]] | ||
| + | |||
'''3 b.''' Visualization with '''showsphere''': | '''3 b.''' Visualization with '''showsphere''': | ||
| Line 121: | Line 122: | ||
* Go ''File'' -> ''Open'', choose '''output_spheres.pdb''' | * Go ''File'' -> ''Open'', choose '''output_spheres.pdb''' | ||
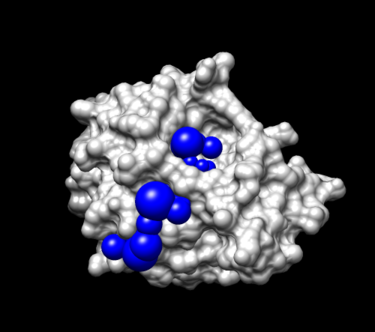
You should see many spheres placed all over the receptor surface. | You should see many spheres placed all over the receptor surface. | ||
| + | |||
| + | [[Image:4tkg.rec_allsph_selectsphere.png|thumb|center|375px|4TKG Receptor surface (light gray) with all spheres (various colors) using the program showsphere for visualization. The Yellow spheres are located in the binding site of the receptor]] | ||
'''Step 4.''' Selecting spheres of interest: | '''Step 4.''' Selecting spheres of interest: | ||
Revision as of 23:18, 2 March 2015
For additional Rizzo Lab tutorials see DOCK Tutorials. Use this link Wiki Formatting as a reference for editing the wiki. This tutorial was developed collaboratively by the AMS 536 class of 2014, using DOCK v6.6.
Contents
I. Introduction
DOCK
DOCK is a molecular docking program used in drug discovery. It was developed by Irwin D. Kuntz, Jr. and colleagues at UCSF (see UCSF DOCK). This program, given a protein binding site and a small molecule, tries to predict the correct binding mode of the small molecule in the binding site, and the associated binding energy. Small molecules with highly favorable binding energies could be new drug leads. This makes DOCK a valuable drug discovery tool. DOCK is typically used to screen massive libraries of millions of compounds against a protein to isolate potential drug leads. These leads are then further studied, and could eventually result in a new, marketable drug. DOCK works well as a screening procedure for generating leads, but is not currently as useful for optimization of those leads.
DOCK 6 uses an incremental construction algorithm called anchor and grow. It is described by a three-step process:
- Rigid portion of ligand (anchor) is docked by geometric methods.
- Non-rigid segments added in layers; energy minimized.
- The resulting configurations are 'pruned' and energy re-minimized, yielding the docked configurations.
Poly ADP Ribose Polymerase (PARP)
Poly (ADP-ribose) polymerase (PARP) is a family of proteins involved in a number of cellular processes involving mainly DNA repair and programmed cell death. (Wikipedia: http://en.wikipedia.org/wiki/Poly_ADP_ribose_polymerase) The particular PARP family member we will focus on is PARP5b (aka: Tankyrase 2) of which the catalytic domains contains 227 amino acid residues. Olaparib (AZD-2281, trade name Lynparza) is an FDA-approved chemotherapeutic agent, developed by KuDOS Pharmaceuticals and later by AstraZeneca. It is an inhibitor of poly ADP ribose polymerase (PARP), an enzyme involved in DNA repair.[1] It acts against cancers in people with hereditary BRCA1 or BRCA2 mutations, which includes many ovarian, breast, and prostate cancers. (Wikipedia: http://en.wikipedia.org/wiki/Olaparib)
In this class, we will perform docking experiments and virtual screening on a crystallographic structure of PARP5b in complex with a small-molecule inhibitor, olaparib (PDB ID: 4TKG).
Organizing Directories
While performing docking, it is convenient to adopt a standard directory structure / naming scheme, so that files are easy to find / identify. For this tutorial, we will use something similar to the following:
~username/AMS536/dock-tutorial/00.files/
/01.dockprep/
/02.surface-spheres/
/03.box-grid/
/04.dock/
/05.mini-virtual-screen/
/06.virtual-screen/
In addition, most of the important files that are derived from the original crystal structure will be given a prefix that is the same as the PDB code, '4TKG'. The following sections in this tutorial will adhere to this directory structure/naming scheme.
II. Preparing the Receptor and Ligand
Matt Elmes
Courtney Singleton
III. Generating Receptor Surface and Spheres
Kellon Belfon
Xingyu
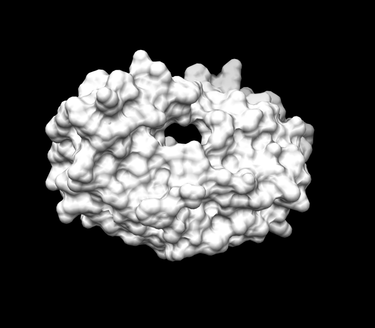
Generating the Receptor Surface
Placing Spheres
Sphgen is a program that generate sets of overlapping spheres that define the shape of a molecule or molecular surface. Spheres are generated over the entire receptor and ligand surface. For further information on how Sphgen generate the spheres for receptors and ligand, please refer to the latest version of the DOCK manual:
<http://dock.compbio.ucsf.edu/DOCK_6/dock6_manual.htm>
To generate spheres using Sphgen follow the steps below:
Step 1. Create an input file name INSPH with the following information:
vim INSPH
4TKG.rec.dms #surface file generated above will be the input file R #flag to place spheres outside (R) or inside (L) of the surface X #flag that informs sphgen of the subset of surface points to be used (X = all points) 0.0 #flag that prevent the generation of large spheres with close surface contacts(default= 0) 4.0 #maximum radius of the spheres generated (default = 4.0 Angstroms) 1.4 #minimum radius of the spheres generated (default = radius of probe) 4TKG.rec.sph #this will be the file which contained the clustered spheres generated
Step 2. Run the program Sphgen using the command sphgen:
sphgen -i INSPH -o OUTSPH
-i is the flag that give sphgen the input file INSPH is INSPH is the file created above that gives sphgen instructions -o is the flag to create the oputput file OUTSPH is the output file with the information of the spheres generated from sphgen
Step 3. Visualization of the spheres generated:
Visualization of the spheres can be done directly with chimera or with the program showsphere
3 a. Visualization directly with Chimera:
- Launch Chimera, choose File -> Open, choose 4TKG.rec.mol2
- choose File -> Open, choose 4TKG.rec.sph
You should have an image like this:
3 b. Visualization with showsphere:
showsphere convert the .sph file into PDB format.
(i) Run showsphere, by typing showsphere into the terminal:
showsphere
You will be prompted with the following questions:
Enter name of sphere cluster file:
4TKG.rec.sph
Enter cluster number to process (<0 = all):
-1
Generate surfaces as well as pdb files (<N>/Y)?
N
Enter name for output file prefix:
output_spheres
Process cluster 0 (contains ALL spheres) (<N>/Y)?
N
-1 is a flag that allow you to see all possible spheres
(ii) Open Chimera
- Launch Chimera, choose File -> Open, choose 4TKG.rec.noH.pdb
- Go File -> Open, choose output_spheres.pdb
You should see many spheres placed all over the receptor surface.
Step 4. Selecting spheres of interest:
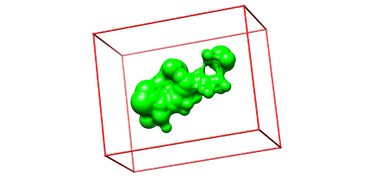
IV. Generating Box and Grid
Box Generation
- Make a new directory and name it: 03.box-grid/
mkdir 03.box-grid
- Make a new file in this directory and name it showbox.in
vim showbox.in
- This will automatically open the file showbox.in. Edit the file showbox.in as follows:
Y # Yes, generate a box 8.0 # Size of the box in Angstroms ../02.surface-spheres/selected_spheres.sph # Sphere.sph file 1 # Cluster number 4TKG.box.pdb # Name of the output file
- Save the file using the command:
:wq
- Run the command:
showbox < showbox.in
Cong Liu
Grid computing
V. Docking a Single Molecule for Pose Reproduction
Michael Cortes
Beibei Zhang
Prajna Shanbhogue
VI. Virtual Screening
george jones
Sam Chiappone