Difference between revisions of "2020 AMBER tutorial with PDBID 3VJK"
Stonybrook (talk | contribs) (→Generating Parameters for the simmulation) |
Stonybrook (talk | contribs) (→Build system with TLeap) |
||
| (47 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
In this tutorial, we will be modeling the dynamics of the ligand with the receptor using AMBER 16. Amber is a molecular dynamics simulation software package. | In this tutorial, we will be modeling the dynamics of the ligand with the receptor using AMBER 16. Amber is a molecular dynamics simulation software package. | ||
| − | + | ||
| + | = Generating Parameters for the simulation = | ||
| + | |||
| + | In order to utilize Amber for molecular dynamic, parameters for the bio molecules will be needed. Luckily, there have been years of parameter development so parameters for the protein do not have to worried about. However, the small ligand does not have parameters in the standard protein force field. Consequently, we will need to generate a fcmod file specific for the ligand. | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | A NOTE FROM JOHN BICKEL: | ||
| + | If you are at this stage, it is recommended to download a fresh PDB from RCSB and remove everything but the protein, and use that as your receptor. The method as outlined below causes some issues that downloading a fresh PDB remedies. | ||
| + | |||
| + | Your receptor, when you give it to tleap, should have NO HYDROGENS and be UNCHARGED. That will be handled by tleap and whatever forcefield you're loading in. | ||
| + | |||
| + | There ARE ways to do what the below aims to do, but using tleap is not the recommended method for doing it. I am keeping it to review it later, but for now you should do the above and continue down to where 000.parameters is made. | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | We will need to have some structures for running the simulation. This can be all stored in a directory called zzz.master. In this directory we will have the following files: | ||
| + | 3vjkFH.pdb 3VJK_hydrogen_protein.mol2 3VJK_ligand_hydrogens.mol2 | ||
| + | Please note that 3VJK_hydrogen_protein.mol2 had to be converted to a pdb using tleap because there were incapability issues that arose in a later step. To convert you do the following: | ||
| + | tleap | ||
| + | rec= loadmol2 3VJK_hydrogen_protein.mol2 | ||
| + | savepdb rec 3vjkFH.pdb | ||
| + | quit | ||
| + | |||
| + | |||
| + | First we will make a specific directory dedicated for the generation of the parameters for the ligand: | ||
| + | mkdir 000.parameters | ||
| + | In this directory we will run the following command: | ||
| + | antechamber -i ./../zzz.master/3VJK_ligand_hydrogens.mol2 -fi mol2 -o 3vjk_ligand_antechamber.mol2 -fo mol2 -at gaff2 -c bcc -rn LIG -nc 2 | ||
| + | notice that will be using gaff2. This stands for general amber force field 2. This will allow us to parameterize ligands for simulations. Additionally, the ligand has a charge of +2 and that was noted with the -nc flag. | ||
| + | Once this command has run, it is beneficial to check to find if any parameters are missing: | ||
| + | parmchk2 -i 3vjk_ligand_antechamber.mol2 -f mol2 -o 3vjk_ligand.am1bcc.frcmod | ||
| + | The output shows that the parameters that were generated were sufficient to continue. | ||
| + | |||
| + | =Build system with TLeap= | ||
| + | Before we can simulate the protein and ligand complex, we must build the whole system together. This involves adding solvent and solvent ions to the protein and ligand complex. In order to accomplish this we will be using tleap. | ||
| + | |||
| + | We will make a new directory for the system: | ||
| + | mkdir 001.tleap_build | ||
| + | now, we will make a tleap.in file--which will contain information about building the system: | ||
| + | vim tleap.in | ||
| + | #!/usr/bin/sh | ||
| + | |||
| + | ###load protein force field | ||
| + | source leaprc.protein.ff14SB | ||
| + | ###load GAFF force field (for our ligand) | ||
| + | source leaprc.gaff | ||
| + | ###load TIP3P (water) force field | ||
| + | source leaprc.water.tip3p | ||
| + | ###load ions frcmod for the tip3p model | ||
| + | loadamberparams frcmod.ionsjc_tip3p | ||
| + | ###needed so we can use igb=8 model | ||
| + | set default PBradii mbondi3 | ||
| + | |||
| + | |||
| + | ###load protein pdb file | ||
| + | rec=loadpdb ./../zzz.master/3vjkFH.pdb | ||
| + | ##@make disulfide bonds | ||
| + | bond rec.328.SG rec.339.SG | ||
| + | bond rec.385.SG rec.394.SG | ||
| + | bond rec.444.SG rec.447.SG | ||
| + | bond rec.454.SG rec.472.SG | ||
| + | bond rec.649.SG rec.762.SG | ||
| + | ###load ligand frcmod/mol2 | ||
| + | loadamberparams ./../000.parameters/3vjk_ligand.am1bcc.frcmod | ||
| + | lig=loadmol2 ./../000.parameters/3vjk_ligand_antechamber.mol2 | ||
| + | |||
| + | ###create gase-phase complex | ||
| + | gascomplex= combine {rec lig} | ||
| + | |||
| + | ###write gas-phase pdb | ||
| + | savepdb gascomplex 3vjk.gas.complex.pdb | ||
| + | ###write gase-phase toplogy and coord files for MMGBSA calc | ||
| + | saveamberparm gascomplex 3vjk.complex.parm7 3vjk.gas.complex.rst7 | ||
| + | saveamberparm rec 3vjk.gas.receptor.parm7 3vjk.gas.receptor.rst7 | ||
| + | saveamberparm lig 3vjk.gas.ligand.parm7 3vjk.gas.ligand.rst7 | ||
| + | |||
| + | ###create solvated complex (albeit redundant) | ||
| + | solvcomplex= combine {rec lig} | ||
| + | |||
| + | ###solvate the system | ||
| + | solvateoct solvcomplex TIP3PBOX 12.0 | ||
| + | |||
| + | ###Neutralize system | ||
| + | addions solvcomplex Cl- 0 | ||
| + | addions solvcomplex Na+ 0 | ||
| + | |||
| + | #write solvated pdb file | ||
| + | savepdb solvcomplex 3vjk.wet.complex.pdb | ||
| + | |||
| + | ###check the system | ||
| + | charge solvcomplex | ||
| + | check solvcomplex | ||
| + | |||
| + | ###write solvated toplogy and coordinate file | ||
| + | saveamberparm solvcomplex 3vjk.wet.complex.parm7 3vjk.wet.complex.rst7 | ||
| + | quit | ||
| + | |||
| + | We now run with tleap -f tleap.in | ||
| + | Note that the number of ions of Cl- and Na+ was set to zero. In this situation, we allowed tleap to use a grid to calculate the ions that are needed to neutralize the system. Later on, we will see that amber correctly added the about of ions to set the system equal to zero. This is an important condition to look at. The system can crash if charges are not resolved. The following files should have been created: | ||
| + | 3vjk.complex.parm7 3vjk.gas.complex.pdb 3vjk.gas.complex.rst7 3vjk.gas.ligand.parm7 3vjk.gas.ligand.rst7 3vjk.gas.receptor.parm7 3vjk.gas.receptor.rst7 3vjk.wet.complex.parm7 3vjk.wet.complex.pdb 3vjk.wet.complex.rst7 | ||
| + | |||
| + | Now that the system is built, it is important to visualize the system to make sure that everything was generated correctly. You may do this using chimera or VMD. Please note that chimera has a special way of loading parm7 and rst7 files. You must go to trajectory and select the two files in order to visualize. Below are the images that were generated using VMD | ||
| + | |||
| + | [[File:3vjk_wet_10.jpg|thumb|center|1400px|The image shows the complete solvated complex with ions]] | ||
| + | |||
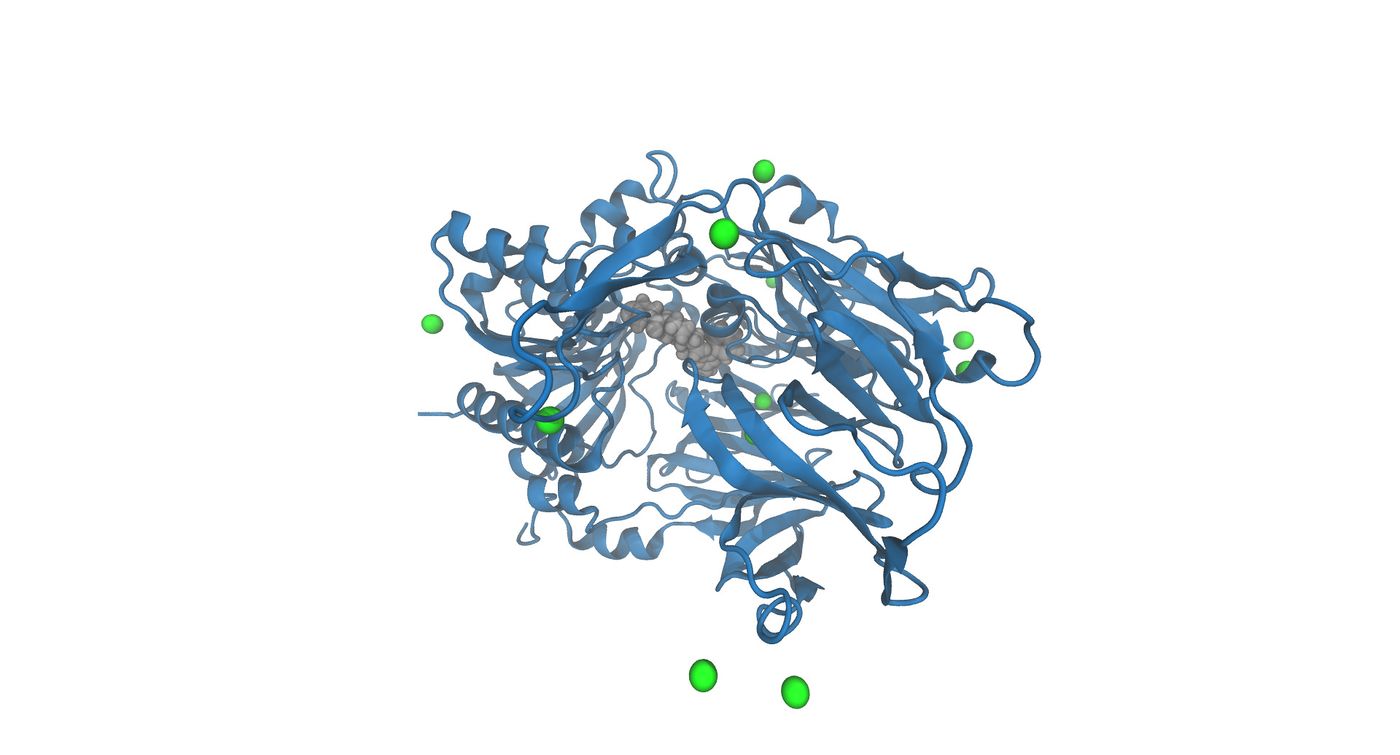
| + | [[File:3vjk_solvated_wet8.jpg|thumb|center|1400px|The image shows the solvated complex without waters. The gray colored object is the ligand in the active site and the green spheres are the Na+ ions]] | ||
| + | |||
| + | =Minimization and Equilibration= | ||
| + | Before we can make production runs, we must minimize and equilibrate the system. This is required because the structure was taken from a crystal. This means that there could be unenergetnyicall favored angles, bonds, clash, ect. We will seek to reduce these un-energetically favored states by relaxing the structure. Additionally, this process is important because new atoms were added to the structure--in particular hydrogens. These hydrogens were added using chimera and may not be in the optimal position. | ||
| + | |||
| + | In order to start this process, we must generate the input files for pmemd | ||
| + | |||
| + | vi 01.min.mdin | ||
| + | |||
| + | Minimize all the hydrogens | ||
| + | &cntrl | ||
| + | imin=1, ! Minimize the initial structure | ||
| + | ntmin=2, ! Use steepest descent Ryota Added | ||
| + | maxcyc=5000, ! Maximum number of cycles for minimization | ||
| + | ntb=1, ! Constant volume | ||
| + | ntp=0, ! No pressure scaling | ||
| + | ntf=1, ! Complete force evaluation | ||
| + | ntwx= 1000, ! Write to trajectory file every ntwx steps | ||
| + | ntpr= 1000, ! Print to mdout every ntpr steps | ||
| + | ntwr= 1000, ! Write a restart file every ntwr steps | ||
| + | cut= 8.0, ! Nonbonded cutoff in Angstroms | ||
| + | ntr=1, ! Turn on restraints | ||
| + | restraintmask="!@H=", ! atoms to be restrained | ||
| + | restraint_wt=5.0, ! force constant for restraint | ||
| + | ntxo=1, ! Write coordinate file in ASCII format | ||
| + | ioutfm=0, ! Write trajectory file in ASCII format | ||
| + | / | ||
| + | |||
| + | 02.equil.mdin | ||
| + | |||
| + | MD simulation | ||
| + | &cntrl | ||
| + | imin=0, ! Perform MD | ||
| + | nstlim=50000 ! Number of MD steps | ||
| + | ntb=2, ! Constant Pressure | ||
| + | ntc=1, ! No SHAKE on bonds between hydrogens | ||
| + | dt=0.001, ! Timestep (ps) | ||
| + | ntp=1, ! Isotropic pressure scaling | ||
| + | barostat=1 ! Berendsen | ||
| + | taup=0.5 ! Pressure relaxtion time (ps) | ||
| + | ntf=1, ! Complete force evaluation | ||
| + | ntt=3, ! Langevin thermostat | ||
| + | gamma_ln=2.0 ! Collision Frequency for thermostat | ||
| + | ig=-1, ! Random seed for thermostat | ||
| + | temp0=298.15 ! Simulation temperature (K) | ||
| + | ntwx= 1000, ! Write to trajectory file every ntwx steps | ||
| + | ntpr= 1000, ! Print to mdout every ntpr steps | ||
| + | ntwr= 1000, ! Write a restart file every ntwr steps | ||
| + | cut= 8.0, ! Nonbonded cutoff in Angstroms | ||
| + | ntr=1, ! Turn on restraints | ||
| + | restraintmask=":!@H=", ! atoms to be restrained | ||
| + | restraint_wt=5.0, ! force constant for restraint | ||
| + | ntxo=1, ! Write coordinate file in ASCII format | ||
| + | ioutfm=0, ! Write trajectory file in ASCII format | ||
| + | iwrap=1, ! iwrap is turned on | ||
| + | / | ||
| + | |||
| + | 03.min.mdin | ||
| + | |||
| + | Minimize all the hydrogens | ||
| + | &cntrl | ||
| + | imin=1, ! Minimize the initial structure | ||
| + | maxcyc=1000, ! Maximum number of cycles for minimization | ||
| + | ntb=1, ! Constant volume | ||
| + | ntp=0, ! No pressure scaling | ||
| + | ntf=1, ! Complete force evaluation | ||
| + | ntwx= 1000, ! Write to trajectory file every ntwx steps | ||
| + | ntpr= 1000, ! Print to mdout every ntpr steps | ||
| + | ntwr= 1000, ! Write a restart file every ntwr steps | ||
| + | cut= 8.0, ! Nonbonded cutoff in Angstroms | ||
| + | ntr=1, ! Turn on restraints | ||
| + | restraintmask="!@H=", ! atoms to be restrained | ||
| + | restraint_wt=2.0, ! force constant for restraint | ||
| + | ntxo=1, ! Write coordinate file in ASCII format | ||
| + | ioutfm=0, ! Write trajectory file in ASCII format | ||
| + | / | ||
| + | |||
| + | 04.min.mdin | ||
| + | Minimize all the hydrogens | ||
| + | &cntrl | ||
| + | imin=1, ! Minimize the initial structure | ||
| + | maxcyc=1000, ! Maximum number of cycles for minimization | ||
| + | ntb=1, ! Constant volume | ||
| + | ntp=0, ! No pressure scaling | ||
| + | ntf=1, ! Complete force evaluation | ||
| + | ntwx= 1000, ! Write to trajectory file every ntwx steps | ||
| + | ntpr= 1000, ! Print to mdout every ntpr steps | ||
| + | ntwr= 1000, ! Write a restart file every ntwr steps | ||
| + | cut= 8.0, ! Nonbonded cutoff in Angstroms | ||
| + | ntr=1, ! Turn on restraints | ||
| + | restraintmask="!@H=", ! atoms to be restrained | ||
| + | restraint_wt=0.1, ! force constant for restraint | ||
| + | ntxo=1, ! Write coordinate file in ASCII format | ||
| + | ioutfm=0, ! Write trajectory file in ASCII format | ||
| + | / | ||
| + | |||
| + | 05.min.mdin | ||
| + | Minimize all the hydrogens | ||
| + | &cntrl | ||
| + | imin=1, ! Minimize the initial structure | ||
| + | maxcyc=1000, ! Maximum number of cycles for minimization | ||
| + | ntb=1, ! Constant volume | ||
| + | ntp=0, ! No pressure scaling | ||
| + | ntf=1, ! Complete force evaluation | ||
| + | ntwx= 1000, ! Write to trajectory file every ntwx steps | ||
| + | ntpr= 1000, ! Print to mdout every ntpr steps | ||
| + | ntwr= 1000, ! Write a restart file every ntwr steps | ||
| + | cut= 8.0, ! Nonbonded cutoff in Angstroms | ||
| + | ntr=1, ! Turn on restraints | ||
| + | restraintmask="!@H=", ! atoms to be restrained | ||
| + | restraint_wt=0.05, ! force constant for restraint | ||
| + | ntxo=1, ! Write coordinate file in ASCII format | ||
| + | ioutfm=0, ! Write trajectory file in ASCII format | ||
| + | / | ||
| + | 06.equil.mdin | ||
| + | MD simulation | ||
| + | &cntrl | ||
| + | imin=0, ! Perform MD | ||
| + | nstlim=50000 ! Number of MD steps | ||
| + | ntb=2, ! Constant Pressure | ||
| + | ntc=1, ! No SHAKE on bonds between hydrogens | ||
| + | dt=0.001, ! Timestep (ps) | ||
| + | ntp=1, ! Isotropic pressure scaling | ||
| + | barostat=1 ! Berendsen | ||
| + | taup=0.5 ! Pressure relaxtion time (ps) | ||
| + | ntf=1, ! Complete force evaluation | ||
| + | ntt=3, ! Langevin thermostat | ||
| + | gamma_ln=2.0 ! Collision Frequency for thermostat | ||
| + | ig=-1, ! Random seed for thermostat | ||
| + | temp0=298.15 ! Simulation temperature (K) | ||
| + | ntwx= 1000, ! Write to trajectory file every ntwx steps | ||
| + | ntpr= 1000, ! Print to mdout every ntpr steps | ||
| + | ntwr= 1000, ! Write a restart file every ntwr steps | ||
| + | cut= 8.0, ! Nonbonded cutoff in Angstroms | ||
| + | ntr=1, ! Turn on restraints | ||
| + | restraintmask="!@H=", ! atoms to be restrained | ||
| + | restraint_wt=1.0, ! force constant for restraint | ||
| + | ntxo=1, ! Write coordinate file in ASCII format | ||
| + | ioutfm=0, ! Write trajectory file in ASCII format | ||
| + | iwrap=1, ! iwrap is turned on | ||
| + | / | ||
| + | 07.equil.mdin | ||
| + | |||
| + | MD simulation | ||
| + | &cntrl | ||
| + | imin=0, ! Perform MD | ||
| + | nstlim=50000 ! Number of MD steps | ||
| + | ntx=5, ! Positions and velocities read formatted | ||
| + | irest=1, ! Restart calculation | ||
| + | ntc=1, ! No SHAKE on for bonds with hydrogen | ||
| + | dt=0.001, ! Timestep (ps) | ||
| + | ntb=2, ! Constant Pressure | ||
| + | ntp=1, ! Isotropic pressure scaling | ||
| + | barostat=1 ! Berendsen | ||
| + | taup=0.5 ! Pressure relaxtion time (ps) | ||
| + | ntf=1, ! Complete force evaluation | ||
| + | ntt=3, ! Langevin thermostat | ||
| + | gamma_ln=2.0 ! Collision Frequency for thermostat | ||
| + | ig=-1, ! Random seed for thermostat | ||
| + | temp0=298.15 ! Simulation temperature (K) | ||
| + | ntwx= 1000, ! Write to trajectory file every ntwx steps | ||
| + | ntpr= 1000, ! Print to mdout every ntpr steps | ||
| + | ntwr= 1000, ! Write a restart file every ntwr steps | ||
| + | cut= 8.0, ! Nonbonded cutoff in Angstroms | ||
| + | ntr=1, ! Turn on restraints | ||
| + | restraintmask="!@H=", ! atoms to be restrained | ||
| + | restraint_wt=0.5, ! force constant for restraint | ||
| + | ntxo=1, ! Write coordinate file in ASCII format | ||
| + | ioutfm=0, ! Write trajectory file in ASCII format | ||
| + | iwrap=1, ! iwrap is turned on | ||
| + | / | ||
| + | 08.equil.mdin | ||
| + | MD simulations | ||
| + | &cntrl | ||
| + | imin=0, ! Perform MD | ||
| + | nstlim=50000 ! Number of MD steps | ||
| + | ntx=5, ! Positions and velocities read formatted | ||
| + | irest=1, ! Restart calculation | ||
| + | ntc=1, ! No SHAKE on for bonds with hydrogen | ||
| + | dt=0.001, ! Timestep (ps) | ||
| + | ntb=2, ! Constant Pressure | ||
| + | ntp=1, ! Isotropic pressure scaling | ||
| + | barostat=1 ! Berendsen | ||
| + | taup=0.5 ! Pressure relaxtion time (ps) | ||
| + | ntf=1, ! Complete force evaluation | ||
| + | ntt=3, ! Langevin thermostat | ||
| + | gamma_ln=2.0 ! Collision Frequency for thermostat | ||
| + | ig=-1, ! Random seed for thermostat | ||
| + | temp0=298.15 ! Simulation temperature (K) | ||
| + | ntwx= 1000, ! Write to trajectory file every ntwx steps | ||
| + | ntpr= 1000, ! Print to mdout every ntpr steps | ||
| + | ntwr= 1000, ! Write a restart file every ntwr steps | ||
| + | cut= 8.0, ! Nonbonded cutoff in Angstroms | ||
| + | ntr=1, ! Turn on restraints | ||
| + | restraintmask=":1-730@CA,C,N", ! atoms to be restrained | ||
| + | restraint_wt=0.1, ! force constant for restraint | ||
| + | ntxo=1, ! Write coordinate file in ASCII format | ||
| + | ioutfm=0, ! Write trajectory file in ASCII format | ||
| + | iwrap=1, ! iwrap is turned on | ||
| + | / | ||
| + | 09.equil.mdin | ||
| + | MD simulations | ||
| + | &cntrl | ||
| + | imin=0, ! Perform MD | ||
| + | nstlim=50000 ! Number of MD steps | ||
| + | ntx=5, ! Positions and velocities read formatted | ||
| + | irest=1, ! Restart calculation | ||
| + | ntc=1, ! No SHAKE on for bonds with hydrogen | ||
| + | dt=0.001, ! Timestep (ps) | ||
| + | ntb=2, ! Constant Pressure | ||
| + | ntp=1, ! Isotropic pressure scaling | ||
| + | barostat=1 ! Berendsen | ||
| + | taup=0.5 ! Pressure relaxtion time (ps) | ||
| + | ntf=1, ! Complete force evaluation | ||
| + | ntt=3, ! Langevin thermostat | ||
| + | gamma_ln=2.0 ! Collision Frequency for thermostat | ||
| + | ig=-1, ! Random seed for thermostat | ||
| + | temp0=298.15 ! Simulation temperature (K) | ||
| + | ntwx= 1000, ! Write to trajectory file every ntwx steps | ||
| + | ntpr= 1000, ! Print to mdout every ntpr steps | ||
| + | ntwr= 1000, ! Write a restart file every ntwr steps | ||
| + | cut= 8.0, ! Nonbonded cutoff in Angstroms | ||
| + | ntr=1, ! Turn on restraints | ||
| + | restraintmask=":1-730@CA,C,N", ! atoms to be restrained | ||
| + | restraint_wt=0.1, ! force constant for restraint | ||
| + | ntxo=1, ! Write coordinate file in ASCII format | ||
| + | ioutfm=0, ! Write trajectory file in ASCII format | ||
| + | iwrap=1, ! iwrap is turned on | ||
| + | / | ||
| + | |||
| + | Now, we have to create a submission script for this to run: | ||
| + | vim mdequilibration.sh | ||
| + | #!/bin/sh | ||
| + | #SBATCH --job-name=3vjk_equilibration | ||
| + | #SBATCH --ntasks-per-node=40 | ||
| + | #SBATCH --nodes=2 | ||
| + | #SBATCH --time=8:00:00 | ||
| + | #SBATCH -p long-40core | ||
| + | |||
| + | cd $SLURM_SUBMIT_DIR | ||
| + | |||
| + | |||
| + | |||
| + | echo "started Equilibration on 'date' " | ||
| + | do_parallel="mpirun pmemd.MPI" | ||
| + | |||
| + | parm7="./../001.tleap_build/3vjk.wet.complex.parm7" | ||
| + | coords="./../001.tleap_build/3vjk.wet.complex" | ||
| + | |||
| + | MDINPUTS=( 01.min 02.equil 03.min 04.min 05.min 06.equil 07.equil 08.equil 09.equil) | ||
| + | |||
| + | for input in ${MDINPUTS[@]}; do | ||
| + | $do_parallel -O -i ${input}.mdin -o ${input}.mdout -p $parm7 -c ${coords}.rst7 -ref ${coords}.rst7 -x ${input}.trj -inf ${input}.info -r ${input}.rst7 | ||
| + | coords=$input | ||
| + | done | ||
| + | |||
| + | echo " Finished equilibration on 'date' " | ||
| + | we will submit this using the following command: | ||
| + | sbatch mdequilibration.sh | ||
Latest revision as of 15:27, 14 April 2021
In this tutorial, we will be modeling the dynamics of the ligand with the receptor using AMBER 16. Amber is a molecular dynamics simulation software package.
Generating Parameters for the simulation
In order to utilize Amber for molecular dynamic, parameters for the bio molecules will be needed. Luckily, there have been years of parameter development so parameters for the protein do not have to worried about. However, the small ligand does not have parameters in the standard protein force field. Consequently, we will need to generate a fcmod file specific for the ligand.
A NOTE FROM JOHN BICKEL:
If you are at this stage, it is recommended to download a fresh PDB from RCSB and remove everything but the protein, and use that as your receptor. The method as outlined below causes some issues that downloading a fresh PDB remedies.
Your receptor, when you give it to tleap, should have NO HYDROGENS and be UNCHARGED. That will be handled by tleap and whatever forcefield you're loading in.
There ARE ways to do what the below aims to do, but using tleap is not the recommended method for doing it. I am keeping it to review it later, but for now you should do the above and continue down to where 000.parameters is made.
We will need to have some structures for running the simulation. This can be all stored in a directory called zzz.master. In this directory we will have the following files:
3vjkFH.pdb 3VJK_hydrogen_protein.mol2 3VJK_ligand_hydrogens.mol2
Please note that 3VJK_hydrogen_protein.mol2 had to be converted to a pdb using tleap because there were incapability issues that arose in a later step. To convert you do the following:
tleap rec= loadmol2 3VJK_hydrogen_protein.mol2 savepdb rec 3vjkFH.pdb quit
First we will make a specific directory dedicated for the generation of the parameters for the ligand:
mkdir 000.parameters
In this directory we will run the following command:
antechamber -i ./../zzz.master/3VJK_ligand_hydrogens.mol2 -fi mol2 -o 3vjk_ligand_antechamber.mol2 -fo mol2 -at gaff2 -c bcc -rn LIG -nc 2
notice that will be using gaff2. This stands for general amber force field 2. This will allow us to parameterize ligands for simulations. Additionally, the ligand has a charge of +2 and that was noted with the -nc flag. Once this command has run, it is beneficial to check to find if any parameters are missing:
parmchk2 -i 3vjk_ligand_antechamber.mol2 -f mol2 -o 3vjk_ligand.am1bcc.frcmod
The output shows that the parameters that were generated were sufficient to continue.
Build system with TLeap
Before we can simulate the protein and ligand complex, we must build the whole system together. This involves adding solvent and solvent ions to the protein and ligand complex. In order to accomplish this we will be using tleap.
We will make a new directory for the system:
mkdir 001.tleap_build
now, we will make a tleap.in file--which will contain information about building the system: vim tleap.in
#!/usr/bin/sh
###load protein force field
source leaprc.protein.ff14SB
###load GAFF force field (for our ligand)
source leaprc.gaff
###load TIP3P (water) force field
source leaprc.water.tip3p
###load ions frcmod for the tip3p model
loadamberparams frcmod.ionsjc_tip3p
###needed so we can use igb=8 model
set default PBradii mbondi3
###load protein pdb file
rec=loadpdb ./../zzz.master/3vjkFH.pdb
##@make disulfide bonds
bond rec.328.SG rec.339.SG
bond rec.385.SG rec.394.SG
bond rec.444.SG rec.447.SG
bond rec.454.SG rec.472.SG
bond rec.649.SG rec.762.SG
###load ligand frcmod/mol2
loadamberparams ./../000.parameters/3vjk_ligand.am1bcc.frcmod
lig=loadmol2 ./../000.parameters/3vjk_ligand_antechamber.mol2
###create gase-phase complex
gascomplex= combine {rec lig}
###write gas-phase pdb
savepdb gascomplex 3vjk.gas.complex.pdb
###write gase-phase toplogy and coord files for MMGBSA calc
saveamberparm gascomplex 3vjk.complex.parm7 3vjk.gas.complex.rst7
saveamberparm rec 3vjk.gas.receptor.parm7 3vjk.gas.receptor.rst7
saveamberparm lig 3vjk.gas.ligand.parm7 3vjk.gas.ligand.rst7
###create solvated complex (albeit redundant)
solvcomplex= combine {rec lig}
###solvate the system
solvateoct solvcomplex TIP3PBOX 12.0
###Neutralize system
addions solvcomplex Cl- 0
addions solvcomplex Na+ 0
#write solvated pdb file
savepdb solvcomplex 3vjk.wet.complex.pdb
###check the system
charge solvcomplex
check solvcomplex
###write solvated toplogy and coordinate file
saveamberparm solvcomplex 3vjk.wet.complex.parm7 3vjk.wet.complex.rst7
quit
We now run with tleap -f tleap.in Note that the number of ions of Cl- and Na+ was set to zero. In this situation, we allowed tleap to use a grid to calculate the ions that are needed to neutralize the system. Later on, we will see that amber correctly added the about of ions to set the system equal to zero. This is an important condition to look at. The system can crash if charges are not resolved. The following files should have been created:
3vjk.complex.parm7 3vjk.gas.complex.pdb 3vjk.gas.complex.rst7 3vjk.gas.ligand.parm7 3vjk.gas.ligand.rst7 3vjk.gas.receptor.parm7 3vjk.gas.receptor.rst7 3vjk.wet.complex.parm7 3vjk.wet.complex.pdb 3vjk.wet.complex.rst7
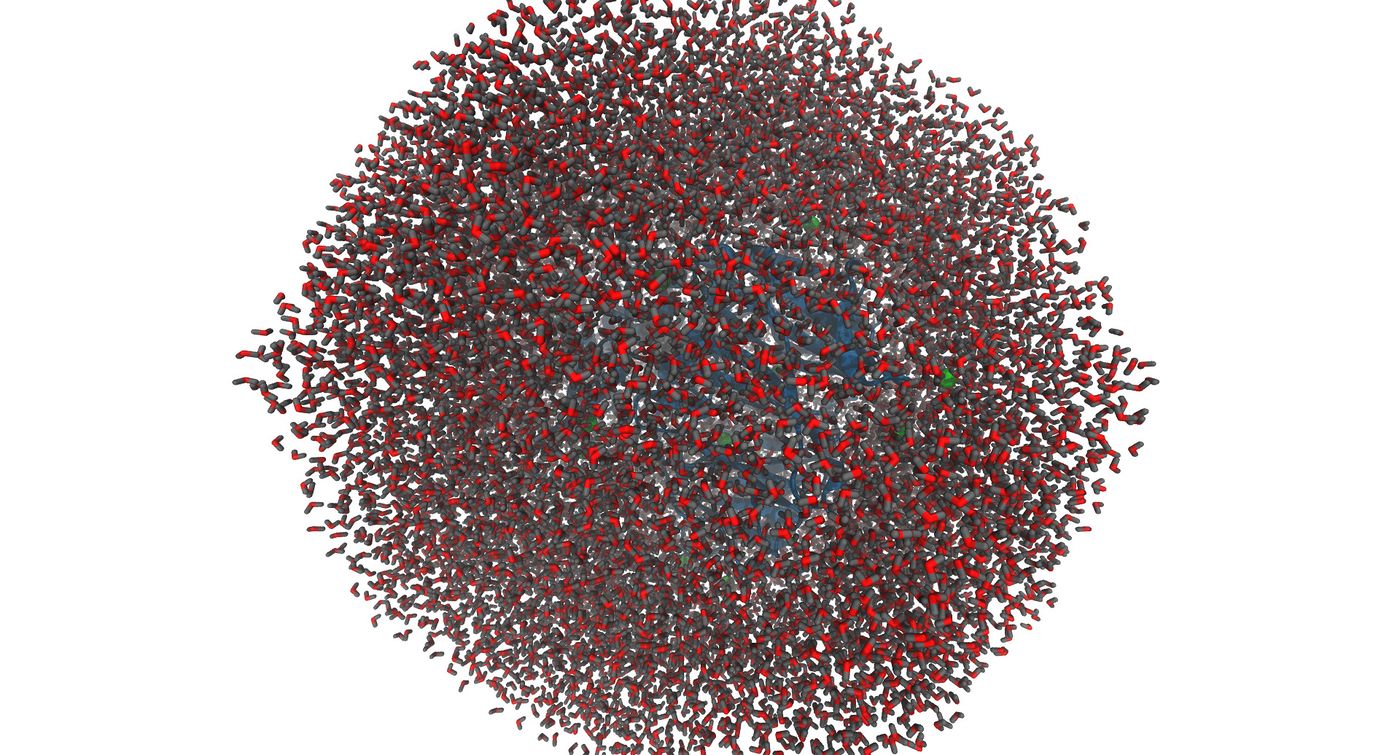
Now that the system is built, it is important to visualize the system to make sure that everything was generated correctly. You may do this using chimera or VMD. Please note that chimera has a special way of loading parm7 and rst7 files. You must go to trajectory and select the two files in order to visualize. Below are the images that were generated using VMD
Minimization and Equilibration
Before we can make production runs, we must minimize and equilibrate the system. This is required because the structure was taken from a crystal. This means that there could be unenergetnyicall favored angles, bonds, clash, ect. We will seek to reduce these un-energetically favored states by relaxing the structure. Additionally, this process is important because new atoms were added to the structure--in particular hydrogens. These hydrogens were added using chimera and may not be in the optimal position.
In order to start this process, we must generate the input files for pmemd
vi 01.min.mdin
Minimize all the hydrogens &cntrl imin=1, ! Minimize the initial structure ntmin=2, ! Use steepest descent Ryota Added maxcyc=5000, ! Maximum number of cycles for minimization ntb=1, ! Constant volume ntp=0, ! No pressure scaling ntf=1, ! Complete force evaluation ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask="!@H=", ! atoms to be restrained restraint_wt=5.0, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format /
02.equil.mdin
MD simulation &cntrl imin=0, ! Perform MD nstlim=50000 ! Number of MD steps ntb=2, ! Constant Pressure ntc=1, ! No SHAKE on bonds between hydrogens dt=0.001, ! Timestep (ps) ntp=1, ! Isotropic pressure scaling barostat=1 ! Berendsen taup=0.5 ! Pressure relaxtion time (ps) ntf=1, ! Complete force evaluation ntt=3, ! Langevin thermostat gamma_ln=2.0 ! Collision Frequency for thermostat ig=-1, ! Random seed for thermostat temp0=298.15 ! Simulation temperature (K) ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask=":!@H=", ! atoms to be restrained restraint_wt=5.0, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format iwrap=1, ! iwrap is turned on /
03.min.mdin
Minimize all the hydrogens &cntrl imin=1, ! Minimize the initial structure maxcyc=1000, ! Maximum number of cycles for minimization ntb=1, ! Constant volume ntp=0, ! No pressure scaling ntf=1, ! Complete force evaluation ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask="!@H=", ! atoms to be restrained restraint_wt=2.0, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format /
04.min.mdin
Minimize all the hydrogens &cntrl imin=1, ! Minimize the initial structure maxcyc=1000, ! Maximum number of cycles for minimization ntb=1, ! Constant volume ntp=0, ! No pressure scaling ntf=1, ! Complete force evaluation ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask="!@H=", ! atoms to be restrained restraint_wt=0.1, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format /
05.min.mdin
Minimize all the hydrogens &cntrl imin=1, ! Minimize the initial structure maxcyc=1000, ! Maximum number of cycles for minimization ntb=1, ! Constant volume ntp=0, ! No pressure scaling ntf=1, ! Complete force evaluation ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask="!@H=", ! atoms to be restrained restraint_wt=0.05, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format /
06.equil.mdin
MD simulation &cntrl imin=0, ! Perform MD nstlim=50000 ! Number of MD steps ntb=2, ! Constant Pressure ntc=1, ! No SHAKE on bonds between hydrogens dt=0.001, ! Timestep (ps) ntp=1, ! Isotropic pressure scaling barostat=1 ! Berendsen taup=0.5 ! Pressure relaxtion time (ps) ntf=1, ! Complete force evaluation ntt=3, ! Langevin thermostat gamma_ln=2.0 ! Collision Frequency for thermostat ig=-1, ! Random seed for thermostat temp0=298.15 ! Simulation temperature (K) ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask="!@H=", ! atoms to be restrained restraint_wt=1.0, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format iwrap=1, ! iwrap is turned on /
07.equil.mdin
MD simulation &cntrl imin=0, ! Perform MD nstlim=50000 ! Number of MD steps ntx=5, ! Positions and velocities read formatted irest=1, ! Restart calculation ntc=1, ! No SHAKE on for bonds with hydrogen dt=0.001, ! Timestep (ps) ntb=2, ! Constant Pressure ntp=1, ! Isotropic pressure scaling barostat=1 ! Berendsen taup=0.5 ! Pressure relaxtion time (ps) ntf=1, ! Complete force evaluation ntt=3, ! Langevin thermostat gamma_ln=2.0 ! Collision Frequency for thermostat ig=-1, ! Random seed for thermostat temp0=298.15 ! Simulation temperature (K) ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask="!@H=", ! atoms to be restrained restraint_wt=0.5, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format iwrap=1, ! iwrap is turned on /
08.equil.mdin
MD simulations &cntrl imin=0, ! Perform MD nstlim=50000 ! Number of MD steps ntx=5, ! Positions and velocities read formatted irest=1, ! Restart calculation ntc=1, ! No SHAKE on for bonds with hydrogen dt=0.001, ! Timestep (ps) ntb=2, ! Constant Pressure ntp=1, ! Isotropic pressure scaling barostat=1 ! Berendsen taup=0.5 ! Pressure relaxtion time (ps) ntf=1, ! Complete force evaluation ntt=3, ! Langevin thermostat gamma_ln=2.0 ! Collision Frequency for thermostat ig=-1, ! Random seed for thermostat temp0=298.15 ! Simulation temperature (K) ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask=":1-730@CA,C,N", ! atoms to be restrained restraint_wt=0.1, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format iwrap=1, ! iwrap is turned on /
09.equil.mdin
MD simulations &cntrl imin=0, ! Perform MD nstlim=50000 ! Number of MD steps ntx=5, ! Positions and velocities read formatted irest=1, ! Restart calculation ntc=1, ! No SHAKE on for bonds with hydrogen dt=0.001, ! Timestep (ps) ntb=2, ! Constant Pressure ntp=1, ! Isotropic pressure scaling barostat=1 ! Berendsen taup=0.5 ! Pressure relaxtion time (ps) ntf=1, ! Complete force evaluation ntt=3, ! Langevin thermostat gamma_ln=2.0 ! Collision Frequency for thermostat ig=-1, ! Random seed for thermostat temp0=298.15 ! Simulation temperature (K) ntwx= 1000, ! Write to trajectory file every ntwx steps ntpr= 1000, ! Print to mdout every ntpr steps ntwr= 1000, ! Write a restart file every ntwr steps cut= 8.0, ! Nonbonded cutoff in Angstroms ntr=1, ! Turn on restraints restraintmask=":1-730@CA,C,N", ! atoms to be restrained restraint_wt=0.1, ! force constant for restraint ntxo=1, ! Write coordinate file in ASCII format ioutfm=0, ! Write trajectory file in ASCII format iwrap=1, ! iwrap is turned on /
Now, we have to create a submission script for this to run: vim mdequilibration.sh
#!/bin/sh
#SBATCH --job-name=3vjk_equilibration
#SBATCH --ntasks-per-node=40
#SBATCH --nodes=2
#SBATCH --time=8:00:00
#SBATCH -p long-40core
cd $SLURM_SUBMIT_DIR
echo "started Equilibration on 'date' "
do_parallel="mpirun pmemd.MPI"
parm7="./../001.tleap_build/3vjk.wet.complex.parm7"
coords="./../001.tleap_build/3vjk.wet.complex"
MDINPUTS=( 01.min 02.equil 03.min 04.min 05.min 06.equil 07.equil 08.equil 09.equil)
for input in ${MDINPUTS[@]}; do
$do_parallel -O -i ${input}.mdin -o ${input}.mdout -p $parm7 -c ${coords}.rst7 -ref ${coords}.rst7 -x ${input}.trj -inf ${input}.info -r ${input}.rst7
coords=$input
done
echo " Finished equilibration on 'date' "
we will submit this using the following command:
sbatch mdequilibration.sh