Difference between revisions of "MD Simulation: Protein in Water"
Stonybrook (talk | contribs) (→GROMACS Coordinate and Topology Files) |
|||
| (53 intermediate revisions by 2 users not shown) | |||
| Line 6: | Line 6: | ||
Ubiquitin is a small and perfectly ordinary protein, making it ideal for an introductory tutorial. A crystal structure can be found at the [http://www.pdb.org PDB] website under the accession code 1UBQ. Before proceeding to the next step, now is a good time to use your favorite text editor to check for a few things: | Ubiquitin is a small and perfectly ordinary protein, making it ideal for an introductory tutorial. A crystal structure can be found at the [http://www.pdb.org PDB] website under the accession code 1UBQ. Before proceeding to the next step, now is a good time to use your favorite text editor to check for a few things: | ||
| − | 1.) Are crystal waters present in the PDB file? In the case of 1UBQ, there are many crystal waters. Ubiquitin is non-enzymatic, though, so | + | 1.) Are crystal waters present in the PDB file? In the case of 1UBQ, there are many crystal waters. Ubiquitin is non-enzymatic, though, so the waters are not important for an active site mechanism, for example. It is a good idea to delete the waters now and allow the GROMACS solvation tool to fill them back in later. |
| − | 2.) Are there any ligands or non-standard residues present in the PDB file? In the case of 1UBQ, there are no ligands or non-standard residues. Ligand preparation and inclusion is covered in another tutorial ([[Protein-Ligand Complex | + | 2.) Are there any ligands or non-standard residues present in the PDB file? In the case of 1UBQ, there are no ligands or non-standard residues. Ligand preparation and inclusion is covered in another tutorial ([[MD Simulation: Protein-Ligand Complex]]). Without going into too much detail, non-standard residues are okay so long as the residue name and atom names conform to the corresponding entry in the residue topology file (.rtp) of the force field you choose. |
| − | 3.) Are there any residues with missing atoms in the PDB file? In the case of 1UBQ, there are not. If there were, however, you would need to take extra preparation steps beforehand to fix the broken residues | + | 3.) Are there any residues with missing atoms in the PDB file? In the case of 1UBQ, there are not. If there were, however, you would need to take extra preparation steps beforehand to fix the broken residues before continuing. |
| Line 25: | Line 25: | ||
occupancy unequal to one (out of 602 atoms). Check your pdb file. | occupancy unequal to one (out of 602 atoms). Check your pdb file. | ||
| − | A quick look back at the original PDB file reveals that the four | + | A quick look back at the original PDB file reveals that the four residues at the C-terminus of ubiquitin in 1UBQ were not as well resolved as the rest of the protein. Pdb2gmx noticed that, too, and adjusted the occupancy of each to '1'. This should be okay for now, as the atomic positions of these residues will be refined later in the minimization and equilibration steps. Aside from that one warning, it appears pdb2gmx changed a few residue names and atom names to conform to the names used in the AMBER03 .rtp file (found within ~/share/gromacs/top/amber03.ff/ of your local GROMACS directory). Ensure that the changes make sense, and it is okay to proceed. |
| Line 52: | Line 52: | ||
GROMACS coordinate files always contain three special lines. The first line is a title - it is good practice to use a detailed title specific to the system being simulated. The second line is the number of atoms. The final line contains the box vectors (in nanometers). The lines in between contain the residue number, residue name, atom name, atom number, and cartesian coordinates (in nanometers) for each atom in the system. | GROMACS coordinate files always contain three special lines. The first line is a title - it is good practice to use a detailed title specific to the system being simulated. The second line is the number of atoms. The final line contains the box vectors (in nanometers). The lines in between contain the residue number, residue name, atom name, atom number, and cartesian coordinates (in nanometers) for each atom in the system. | ||
| + | |||
===Topology file=== | ===Topology file=== | ||
| Line 89: | Line 90: | ||
Protein_chain_A 1 | Protein_chain_A 1 | ||
| − | The first section is an include statement that, when this file is processed, | + | The first section is an include statement that, when this file is processed, pastes the bonded and non-bonded information specific to your force field directly into the topology file. It is a good idea to familiarize yourself with the contents of the 'forcefield.itp' file, and the files included within it, all found under ~/share/gromacs/top/amber03.ff/. |
The second section (usually) contains 6 or 7 sub-sections. First, under '[ moleculetype ]' is the name of the molecule followed by a number. In this case, it is by default called 'Protein_chain_A'. The number <i>N</i> indicates that for this molecule, exclude non-bonded interactions of all bonded neighbors up to <i>N</i> bonds away. Unless you have a very good idea of what you are doing, you should not change this number. A given moleculetype is followed by (and it must be in this order) atom information, bond information, pair-wise exclusion information, angles information, dihedral information, improper dihedral information, and, optionally, position restraint information. Not all of the sections must be present. Further, this entire section is repeated for each type of molecule that you have. For example, if you are simulating a protein in water, you will have two consecutive '[ moleculetype ]' sections. | The second section (usually) contains 6 or 7 sub-sections. First, under '[ moleculetype ]' is the name of the molecule followed by a number. In this case, it is by default called 'Protein_chain_A'. The number <i>N</i> indicates that for this molecule, exclude non-bonded interactions of all bonded neighbors up to <i>N</i> bonds away. Unless you have a very good idea of what you are doing, you should not change this number. A given moleculetype is followed by (and it must be in this order) atom information, bond information, pair-wise exclusion information, angles information, dihedral information, improper dihedral information, and, optionally, position restraint information. Not all of the sections must be present. Further, this entire section is repeated for each type of molecule that you have. For example, if you are simulating a protein in water, you will have two consecutive '[ moleculetype ]' sections. | ||
| Line 102: | Line 103: | ||
[ moleculetype ] | [ moleculetype ] | ||
| − | + | SOL 1 | |
... | ... | ||
| Line 110: | Line 111: | ||
[ molecules ] | [ molecules ] | ||
Protein 1 | Protein 1 | ||
| − | + | SOL 10000 | |
where the moleculetypes under '[ molecules ]' are listed in the same order as they appear in the topology file. Note that the topology file you generated already contains an #include statement to include the moleculetype for TIP3P water and for ions, but because there are no water molecules or ions in our coordinate file, they are not yet listed under '[ molecules ]'. Chapter 5 of the GROMACS manual is an excellent resource for further information regarding molecular topologies. | where the moleculetypes under '[ molecules ]' are listed in the same order as they appear in the topology file. Note that the topology file you generated already contains an #include statement to include the moleculetype for TIP3P water and for ions, but because there are no water molecules or ions in our coordinate file, they are not yet listed under '[ molecules ]'. Chapter 5 of the GROMACS manual is an excellent resource for further information regarding molecular topologies. | ||
| + | |||
===Position restraints file=== | ===Position restraints file=== | ||
| Line 123: | Line 125: | ||
... | ... | ||
| − | Each line indicates an atom number, a functional form (1 = harmonic restraint), and | + | Each line indicates an atom number, a functional form (1 = harmonic restraint), and restoring forces (units of kJ mol<sup>-1</sup> nm<sup>-2</sup>) in cartesian space. By default, only the protein backbone atoms are listed in this file. Looking back at the topology file, you can see that if 'POSRES' is defined when you begin simulation, then these parameters will be included in the molecular topology, thus restraining the backbone of the protein during simulation. If 'POSRES' is not defined, then the position restraints are ignored. |
| + | |||
==Box Preparation== | ==Box Preparation== | ||
| + | In the next few steps, you will define a box size for your system, center the protein in the box, solvate, and finally, add counterions to the system. The files you need to start this step are: | ||
| + | |||
| + | 1.) protein.gro # gromacs-format coordinate file | ||
| + | 2.) topol.top # gromacs-format topology file | ||
| + | 3.) posre.itp # position restraint file | ||
| + | |||
| + | |||
| + | ===editconf=== | ||
| + | The GROMACS tool editconf is very useful to change the format of your coordinate files, to rotate and translate coordinate files, to define the box size, among other things. Typing 'editconf -h' will display a brief description of its capabilities. To create a rectangular box around the protein, type: | ||
| + | |||
| + | editconf -f protein.gro -o protein_box.gro -bt triclinic -d 1.2 -c | ||
| + | |||
| + | The only input needed is the coordinate file generated previously. With '-bt triclinic' you are choosing to create a rectangular box. It is important to note that rectangular boxes can be very inefficient, especially for globular proteins like ubiquitin. In this tutorial, we will keep with the rectangular box, but in the future, consider using '-bt dodecahedron' for globular proteins. The option '-d 1.2' creates a buffer of 1.2 nm between the outside of the protein and the edge of the box, and '-c' centers the protein in the box and puts the corner of the box at {0, 0, 0} in cartesian space. The output file is called 'protein_box.gro'. | ||
| + | |||
| + | The most important question to ask now is <i>why choose 1.2 nm?</i> The short answer is that you don't want the protein to 'see' its periodic image across the boundary of the box. If you refer to the original paper for the AMBER 03 force field (Duan, <i>et al., J. Comput. Chem.</i> <b>2003,</b> <i>24,</i> 1999-2012), they use short-range VDW and electrostatic cut-offs of 0.8 nm - when you begin simulating, it is very important to do the same. In order for the protein to avoid seeing its image across the periodic boundary, it must be at least twice the cut-off distance from the next nearest image of itself. That being said, the space between the protein and the edge of the box only really needs to be slightly larger than 0.8 nm, but it is still a good idea to include extra space especially if there is a chance the protein might unfold a little bit or if is an oblong shape. It is better to have a slightly larger box size now than to find out later that your protein was interacting with its periodic image during the simulation. (See the image below). | ||
| + | |||
| + | |||
| + | <center>[[Image:pbc_overlap.jpg]]<br> | ||
| + | |||
| + | <i>The dashed black squares indicate the box boundaries, and the dashed red circles indicate the short-range VDW and electrostatic <br> | ||
| + | cut-offs. So long as the red circles do not overlap, the protein will not be able to "see" its periodic image.</i></center> | ||
| + | |||
| + | |||
| + | ===genbox=== | ||
| + | The next GROMACS command, 'genbox', will fill the box with solvent molecules. | ||
| + | |||
| + | genbox -cp protein_box.gro -cs spc216.gro -o protein_sol.gro -p topol.top | ||
| + | |||
| + | The '-cp' flag specifies the coordinate file of the system to be solvated, and '-cs spc216.gro' tells genbox to use an SPC (a simple three-point water model) coordinate file to populate the box. Thee output coordinate file is called 'protein_sol.gro'. The topology file, 'topol.top', is also provided on the command line. If you check the end of the topology file, you can see that it has been updated in the following way: | ||
| + | |||
| + | [ molecules ] | ||
| + | Protein_chain_A 1 | ||
| + | SOL 5760 | ||
| + | |||
| + | At this point, it is useful to open 'protein_sol.gro' in VMD to ensure that the protein is correctly placed in the center of the box and there are no strange solvent artifacts. | ||
| + | |||
| + | |||
| + | <center>[[Image:ubq_sol.jpg]]<br> | ||
| + | |||
| + | <i>1UBQ in a triclinic box solvated with 5,760 water molecules.</i></center> | ||
| + | |||
| + | |||
| + | ===genion=== | ||
| + | The final step before simulation is to add enough ions to the system to neutralize the net charge or, alternatively, add enough ions to neutralize the net charge <i>and</i> reach some physiological concentration. If you are simulating a system in an attempt to replicate some experimental observable, for example, it is important to use the same salt concentration in your system as is used in the experiment. A standard value for salt concentration often used to replicate human physiology is 100 mM. | ||
| + | |||
| + | The 'genion' tool searches through your coordinate file and will randomly replace water molecules with ions. In order for it to work, however, it requires a pre-processed input file (with extension .tpr) that contains all of the information from both the coordinate and topology files. In order to generate such a file, you will use the tool 'grompp'. Aside from the coordinate and topology file, 'grompp' also requires that you provide a MD parameter file (.mdp) as input. Because the .tpr file you are generating is not going to be used for dynamics, rather it is just going to be used to add ions, the contents of the .mdp file are not important. Nevertheless, it still must be provided on the command line. An example .mdp file with all of the default parameters can be found here: [[genion.mdp]]. To create the input file, execute: | ||
| + | |||
| + | grompp -f genion.mdp -c protein_sol.gro -p topol.top -o genion_input.tpr | ||
| + | |||
| + | As always, read what was output to the screen, and if there are no major Errors or Warnings, than it is okay to proceed. The net charge of the ubiquitin system is already 0 (a running total of the charge can be found in the '[ atoms ]' section of the protein moleculetype under the definition 'qtot'), so instead of neutralizing the system, add enough NaCl to reach 100 mM salt concentration. To do so, execute: | ||
| + | |||
| + | genion -s genion_input.tpr -p topol.top -o protein_sol_nacl.gro -pname NA -pq 1 -nname CL -nq -1 -conc 0.1 -neutral | ||
| + | |||
| + | It is a good idea to read the output from 'genion -h' in order to gain a full understanding of the command line options. When prompted, choose group 13 (SOL). Genion then chooses 22 solvent molecules and replaces half with sodium ions, and the other half with chloride ions. The topology file is also provided on the command line so that it may be updated accordingly. The '[ molecules ]' section has been modified by removing 22 SOL molecules and adding 11 each of NA and CL molecules. | ||
| + | |||
| + | [ molecules ] | ||
| + | Protein_chain_A 1 | ||
| + | SOL 5738 | ||
| + | NA 11 | ||
| + | CL 11 | ||
| + | |||
| + | |||
| + | <center>[[Image:ubq_ions.jpg]]<br> | ||
| + | |||
| + | <i>1UBQ in a triclinic box solvated with 5,738 water molecules and 22 ions.</i></center> | ||
| + | |||
==Energy Minimization== | ==Energy Minimization== | ||
| + | Prior to running actual dynamics, you will need to perform an energy minimization. The purpose of a minimization is to relax the molecular geometry of the system in order to get rid of any atomic clashes or other irregularities that may exist. The files you need to start this step are: | ||
| + | |||
| + | 1.) protein_sol_nacl.gro # gromacs-format coordinate file | ||
| + | 2.) topol.top # gromacs-format topology file | ||
| + | 3.) posre.itp # position restraint file | ||
| + | 4.) ubq_min.mdp # energy minimization parameter file | ||
| + | |||
| + | The file 'ubq_min.mdp' contains the run parameters for the minimization. A copy of this file can be found here: [[ubq_min.mdp]]. The comments in the file help to explain the purpose of each parameter. However, a review of Chapter 7 in the GROMACS manual would be invaluable (specific sections are listed in the .mdp file). In brief, this parameter file calls for a steepest descent energy minimization not to exceed 2,000 steps. The minimization will be considered successful if the maximum force on any atom is less than 1000 kJ/mol/nm, at which point the minimization will stop. Other parameters of note include periodic boundary conditions in the x-, y-, and z-directions, 0.8 nm cut-offs for short-range VDW and electrostatic interactions, and particle-mesh Ewald long-range electrostatics. If any of these settings do not make sense, it is unwise to proceed without reading about them in Chapter 7 of the manual. | ||
| + | |||
| + | The main integration engine of GROMACS is a tool called 'mdrun'. As input, it requires a pre-processed run input file (.tpr) that contains the system's coordinates, topology, and parameters for the minimization itself. As before, 'grompp' is used to generate that file. Execute the following command: | ||
| + | |||
| + | grompp -f ubq_min.mdp -c protein_sol_nacl.gro -p topol.top -o input_min.tpr | ||
| + | |||
| + | Read the screen output carefully. If the .tpr file is generated successfully, you can perform the minimization: | ||
| + | |||
| + | mdrun -s input_min.tpr -deffnm ubiquitin_min -v | ||
| + | |||
| + | By using the '-v' option, you can watch the minimization progress on the screen. Using the steepest descent method, small systems may equilibrate after only a few hundred steps; larger systems may may take several thousand steps. It mostly depends on the complexity of the system and the quality of the original starting structure. Once the minimization is complete (which should take around 500 steps), you will find that several files have been written to disk, all with the prefix 'ubiquitin_min'. They are as follows: | ||
| + | |||
| + | 1.) ubiquitin_min.gro # system coordinates at the final step | ||
| + | 2.) ubiquitin_min.trr # trajectory file | ||
| + | 3.) ubiquitin_min.log # log file | ||
| + | 4.) ubiquitin_min.edr # energy file | ||
| + | |||
| + | The first file, 'ubiquitin_min.gro', contains the fully minimized coordinates of the system (presuming the minimization was successful). The second file, 'ubiquitin_min.trr' contains a trajectory of the minimization. The frequency with which frames are written to the trajectory is specified in the .mdp file that was passed to grompp. You can view this in VMD by first loading the <i>original</i> coordinate file (protein_sol_nacl.gro), followed by the trajectory file. The log file contains information about the run parameters used for the minimization, as well as various system energies during the minimization. It is convenient to check the progress of long MD simulations by following the end of this file. Finally, ubiquitin_min.edr is a binary energy file that can be read by the GROMACS tool 'g_energy', yielding many pieces of valuable information. In this case, you can use it to check the potential energy of the system as a function of minimization step. Execute the following command: | ||
| + | |||
| + | g_energy -f ubiquitin_min.edr -o energy.xvg | ||
| + | |||
| + | A prompt will appear asking for a specific term you would like to analyze: | ||
| + | |||
| + | Select the terms you want from the following list by | ||
| + | selecting either (part of) the name or the number or a combination. | ||
| + | End your selection with an empty line or a zero. | ||
| + | ------------------------------------------------------------------- | ||
| + | 1 Bond 2 Angle 3 Proper-Dih. 4 Improper-Dih. | ||
| + | 5 LJ-14 6 Coulomb-14 7 LJ-(SR) 8 Disper.-corr. | ||
| + | 9 Coulomb-(SR) 10 Coul.-recip. 11 Potential 12 Pres.-DC | ||
| + | 13 Pressure 14 Vir-XX 15 Vir-XY 16 Vir-XZ | ||
| + | 17 Vir-YX 18 Vir-YY 19 Vir-YZ 20 Vir-ZX | ||
| + | 21 Vir-ZY 22 Vir-ZZ 23 Pres-XX 24 Pres-XY | ||
| + | 25 Pres-XZ 26 Pres-YX 27 Pres-YY 28 Pres-YZ | ||
| + | 29 Pres-ZX 30 Pres-ZY 31 Pres-ZZ 32 #Surf*SurfTen | ||
| + | 33 Mu-X 34 Mu-Y 35 Mu-Z 36 T-rest | ||
| + | |||
| + | Choose '11' for the potential energy, and hit return on an empty line to finish. A file called 'energy.xvg' will be written to file. This is a simple two-column data file that can be plotted in a program such as Grace: | ||
| + | |||
| + | <center>[[Image:ubq_energy.jpg]]<br> | ||
| − | + | <i>Potential energy of system as a function of minimization step.</i></center> | |
| − | + | Convergence of the potential energy in a system to a large negative number (on the order of ~10^5) is a very good indication that the energy minimization was successful. If you find that for your system the potential energy has converged, and if the maximum force on any atom is below a reasonable tolerance cut-off, then it is okay to proceed to Equilibration MD. Otherwise, you may need to use other minimization methods (such as conjugate gradients or L-BFGS) to reach a lower energy well. The implementation of these are described in detail in the GROMACS manual. | |
| − | == | + | ==[[MD Simulation: Protein in Water (Pt. 2)]]== |
Latest revision as of 08:22, 29 February 2012
Contents
Purpose
The purpose of this tutorial is to provide an introduction to the fundamental commands needed to set up, run, and analyze MD simulations in GROMACS. This tutorial assumes you have already correctly installed GROMACS. This tutorial was written using GROMACS 4.5.4.
Download and Prepare PDB File
Ubiquitin is a small and perfectly ordinary protein, making it ideal for an introductory tutorial. A crystal structure can be found at the PDB website under the accession code 1UBQ. Before proceeding to the next step, now is a good time to use your favorite text editor to check for a few things:
1.) Are crystal waters present in the PDB file? In the case of 1UBQ, there are many crystal waters. Ubiquitin is non-enzymatic, though, so the waters are not important for an active site mechanism, for example. It is a good idea to delete the waters now and allow the GROMACS solvation tool to fill them back in later.
2.) Are there any ligands or non-standard residues present in the PDB file? In the case of 1UBQ, there are no ligands or non-standard residues. Ligand preparation and inclusion is covered in another tutorial (MD Simulation: Protein-Ligand Complex). Without going into too much detail, non-standard residues are okay so long as the residue name and atom names conform to the corresponding entry in the residue topology file (.rtp) of the force field you choose.
3.) Are there any residues with missing atoms in the PDB file? In the case of 1UBQ, there are not. If there were, however, you would need to take extra preparation steps beforehand to fix the broken residues before continuing.
GROMACS Coordinate and Topology Files
The first step in MD simulation with GROMACS is to create GROMACS-compatible coordinate (.gro) and topology (.top) files. Strictly speaking, .pdb files are GROMACS-compatible, but we might as well convert to a .gro coordinate file to keep in line with the spirit of the tutorial. The command line execution looks like this:
pdb2gmx -f 1UBQ.pdb -o protein.gro -p topol.top -ignh -v
When prompted, choose '1' for the AMBER03 force field, and '1' again for the TIP3P water model. These selections are fine for this tutorial, but make sure you think very carefully about your choice before picking a force field in your research. You may ask yourself, should I use an all-atom force field or a united-atom force field? Is my force field selection compatible with lipids or small molecules that I want to use? Can I compare the results of simulations using my force field to other simulations I performed previously / found in the literature? etc.
After you have picked the force field and a solvent model compatible with that force field, it is a good idea to read through the output on the screen to make sure there are no Errors or Warnings. In this case, it looks like there actually was one Warning:
WARNING: there were 0 atoms with zero occupancy and 28 atoms with
occupancy unequal to one (out of 602 atoms). Check your pdb file.
A quick look back at the original PDB file reveals that the four residues at the C-terminus of ubiquitin in 1UBQ were not as well resolved as the rest of the protein. Pdb2gmx noticed that, too, and adjusted the occupancy of each to '1'. This should be okay for now, as the atomic positions of these residues will be refined later in the minimization and equilibration steps. Aside from that one warning, it appears pdb2gmx changed a few residue names and atom names to conform to the names used in the AMBER03 .rtp file (found within ~/share/gromacs/top/amber03.ff/ of your local GROMACS directory). Ensure that the changes make sense, and it is okay to proceed.
There are three output files resulting from pdb2gmx:
1.) protein.gro # gromacs-format coordinate file 2.) topol.top # gromacs-format topology file 3.) posre.itp # position restraint file
It is very important to know and understand the contents of each file before continuing. Make sure you look through each file until you are able to make sense of the information contained within each.
Coordinate file
The GROMACS coordinate file contains the following:
Ubiquitin from 1UBQ - no water molecules (yet)
1231
1MET N 1 2.734 2.443 0.261
1MET H1 2 2.807 2.485 0.207
1MET H2 3 2.771 2.413 0.349
...
76GLY C 1229 4.003 3.999 3.543
76GLY OC1 1230 4.086 3.957 3.625
76GLY OC2 1231 3.893 4.052 3.569
3.06857 3.23738 3.66170
GROMACS coordinate files always contain three special lines. The first line is a title - it is good practice to use a detailed title specific to the system being simulated. The second line is the number of atoms. The final line contains the box vectors (in nanometers). The lines in between contain the residue number, residue name, atom name, atom number, and cartesian coordinates (in nanometers) for each atom in the system.
Topology file
The GROMACS topology file contains the following sections:
#include "amber03.ff/forcefield.itp"
[ moleculetype ] Protein_chain_A 3 [ atoms ] ... [ bonds ] ... [ pairs ] ... [ angles ] ... [ dihedrals ] ... [ dihedrals ] ... #ifdef POSRES #include "posre.itp" #endif
[ system ] Ubiquitin [ molecules ] Protein_chain_A 1
The first section is an include statement that, when this file is processed, pastes the bonded and non-bonded information specific to your force field directly into the topology file. It is a good idea to familiarize yourself with the contents of the 'forcefield.itp' file, and the files included within it, all found under ~/share/gromacs/top/amber03.ff/.
The second section (usually) contains 6 or 7 sub-sections. First, under '[ moleculetype ]' is the name of the molecule followed by a number. In this case, it is by default called 'Protein_chain_A'. The number N indicates that for this molecule, exclude non-bonded interactions of all bonded neighbors up to N bonds away. Unless you have a very good idea of what you are doing, you should not change this number. A given moleculetype is followed by (and it must be in this order) atom information, bond information, pair-wise exclusion information, angles information, dihedral information, improper dihedral information, and, optionally, position restraint information. Not all of the sections must be present. Further, this entire section is repeated for each type of molecule that you have. For example, if you are simulating a protein in water, you will have two consecutive '[ moleculetype ]' sections.
The final section only includes two sub-sections. Under '[ system ]' is a system title chosen by the user, and under '[ molecules ]' is a list of the moleculetypes found in the topology file, followed by the number of times each moleculetype appears in the coordinate file. For example if you have a coordinate file that contains a protein in 10,000 water molecules, your topology file should look roughly like:
#include "amber03.ff/forcefield.itp"
[ moleculetype ] Protein 3 ...
[ moleculetype ] SOL 1 ...
[ system ] Protein in water [ molecules ] Protein 1 SOL 10000
where the moleculetypes under '[ molecules ]' are listed in the same order as they appear in the topology file. Note that the topology file you generated already contains an #include statement to include the moleculetype for TIP3P water and for ions, but because there are no water molecules or ions in our coordinate file, they are not yet listed under '[ molecules ]'. Chapter 5 of the GROMACS manual is an excellent resource for further information regarding molecular topologies.
Position restraints file
The final file that was created by pdb2gmx is the position restraints file:
[ position_restraints ]
1 1 1000 1000 1000
5 1 1000 1000 1000
7 1 1000 1000 1000
...
Each line indicates an atom number, a functional form (1 = harmonic restraint), and restoring forces (units of kJ mol-1 nm-2) in cartesian space. By default, only the protein backbone atoms are listed in this file. Looking back at the topology file, you can see that if 'POSRES' is defined when you begin simulation, then these parameters will be included in the molecular topology, thus restraining the backbone of the protein during simulation. If 'POSRES' is not defined, then the position restraints are ignored.
Box Preparation
In the next few steps, you will define a box size for your system, center the protein in the box, solvate, and finally, add counterions to the system. The files you need to start this step are:
1.) protein.gro # gromacs-format coordinate file 2.) topol.top # gromacs-format topology file 3.) posre.itp # position restraint file
editconf
The GROMACS tool editconf is very useful to change the format of your coordinate files, to rotate and translate coordinate files, to define the box size, among other things. Typing 'editconf -h' will display a brief description of its capabilities. To create a rectangular box around the protein, type:
editconf -f protein.gro -o protein_box.gro -bt triclinic -d 1.2 -c
The only input needed is the coordinate file generated previously. With '-bt triclinic' you are choosing to create a rectangular box. It is important to note that rectangular boxes can be very inefficient, especially for globular proteins like ubiquitin. In this tutorial, we will keep with the rectangular box, but in the future, consider using '-bt dodecahedron' for globular proteins. The option '-d 1.2' creates a buffer of 1.2 nm between the outside of the protein and the edge of the box, and '-c' centers the protein in the box and puts the corner of the box at {0, 0, 0} in cartesian space. The output file is called 'protein_box.gro'.
The most important question to ask now is why choose 1.2 nm? The short answer is that you don't want the protein to 'see' its periodic image across the boundary of the box. If you refer to the original paper for the AMBER 03 force field (Duan, et al., J. Comput. Chem. 2003, 24, 1999-2012), they use short-range VDW and electrostatic cut-offs of 0.8 nm - when you begin simulating, it is very important to do the same. In order for the protein to avoid seeing its image across the periodic boundary, it must be at least twice the cut-off distance from the next nearest image of itself. That being said, the space between the protein and the edge of the box only really needs to be slightly larger than 0.8 nm, but it is still a good idea to include extra space especially if there is a chance the protein might unfold a little bit or if is an oblong shape. It is better to have a slightly larger box size now than to find out later that your protein was interacting with its periodic image during the simulation. (See the image below).
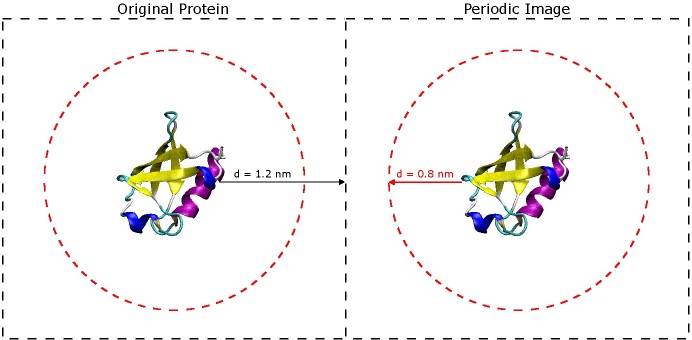
The dashed black squares indicate the box boundaries, and the dashed red circles indicate the short-range VDW and electrostatic
genbox
The next GROMACS command, 'genbox', will fill the box with solvent molecules.
genbox -cp protein_box.gro -cs spc216.gro -o protein_sol.gro -p topol.top
The '-cp' flag specifies the coordinate file of the system to be solvated, and '-cs spc216.gro' tells genbox to use an SPC (a simple three-point water model) coordinate file to populate the box. Thee output coordinate file is called 'protein_sol.gro'. The topology file, 'topol.top', is also provided on the command line. If you check the end of the topology file, you can see that it has been updated in the following way:
[ molecules ] Protein_chain_A 1 SOL 5760
At this point, it is useful to open 'protein_sol.gro' in VMD to ensure that the protein is correctly placed in the center of the box and there are no strange solvent artifacts.
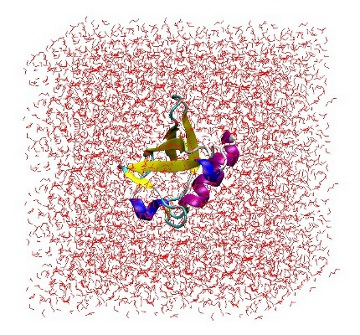
1UBQ in a triclinic box solvated with 5,760 water molecules.
genion
The final step before simulation is to add enough ions to the system to neutralize the net charge or, alternatively, add enough ions to neutralize the net charge and reach some physiological concentration. If you are simulating a system in an attempt to replicate some experimental observable, for example, it is important to use the same salt concentration in your system as is used in the experiment. A standard value for salt concentration often used to replicate human physiology is 100 mM.
The 'genion' tool searches through your coordinate file and will randomly replace water molecules with ions. In order for it to work, however, it requires a pre-processed input file (with extension .tpr) that contains all of the information from both the coordinate and topology files. In order to generate such a file, you will use the tool 'grompp'. Aside from the coordinate and topology file, 'grompp' also requires that you provide a MD parameter file (.mdp) as input. Because the .tpr file you are generating is not going to be used for dynamics, rather it is just going to be used to add ions, the contents of the .mdp file are not important. Nevertheless, it still must be provided on the command line. An example .mdp file with all of the default parameters can be found here: genion.mdp. To create the input file, execute:
grompp -f genion.mdp -c protein_sol.gro -p topol.top -o genion_input.tpr
As always, read what was output to the screen, and if there are no major Errors or Warnings, than it is okay to proceed. The net charge of the ubiquitin system is already 0 (a running total of the charge can be found in the '[ atoms ]' section of the protein moleculetype under the definition 'qtot'), so instead of neutralizing the system, add enough NaCl to reach 100 mM salt concentration. To do so, execute:
genion -s genion_input.tpr -p topol.top -o protein_sol_nacl.gro -pname NA -pq 1 -nname CL -nq -1 -conc 0.1 -neutral
It is a good idea to read the output from 'genion -h' in order to gain a full understanding of the command line options. When prompted, choose group 13 (SOL). Genion then chooses 22 solvent molecules and replaces half with sodium ions, and the other half with chloride ions. The topology file is also provided on the command line so that it may be updated accordingly. The '[ molecules ]' section has been modified by removing 22 SOL molecules and adding 11 each of NA and CL molecules.
[ molecules ] Protein_chain_A 1 SOL 5738 NA 11 CL 11
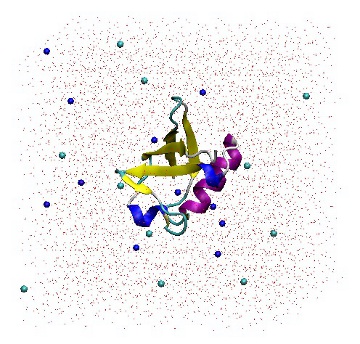
1UBQ in a triclinic box solvated with 5,738 water molecules and 22 ions.
Energy Minimization
Prior to running actual dynamics, you will need to perform an energy minimization. The purpose of a minimization is to relax the molecular geometry of the system in order to get rid of any atomic clashes or other irregularities that may exist. The files you need to start this step are:
1.) protein_sol_nacl.gro # gromacs-format coordinate file 2.) topol.top # gromacs-format topology file 3.) posre.itp # position restraint file 4.) ubq_min.mdp # energy minimization parameter file
The file 'ubq_min.mdp' contains the run parameters for the minimization. A copy of this file can be found here: ubq_min.mdp. The comments in the file help to explain the purpose of each parameter. However, a review of Chapter 7 in the GROMACS manual would be invaluable (specific sections are listed in the .mdp file). In brief, this parameter file calls for a steepest descent energy minimization not to exceed 2,000 steps. The minimization will be considered successful if the maximum force on any atom is less than 1000 kJ/mol/nm, at which point the minimization will stop. Other parameters of note include periodic boundary conditions in the x-, y-, and z-directions, 0.8 nm cut-offs for short-range VDW and electrostatic interactions, and particle-mesh Ewald long-range electrostatics. If any of these settings do not make sense, it is unwise to proceed without reading about them in Chapter 7 of the manual.
The main integration engine of GROMACS is a tool called 'mdrun'. As input, it requires a pre-processed run input file (.tpr) that contains the system's coordinates, topology, and parameters for the minimization itself. As before, 'grompp' is used to generate that file. Execute the following command:
grompp -f ubq_min.mdp -c protein_sol_nacl.gro -p topol.top -o input_min.tpr
Read the screen output carefully. If the .tpr file is generated successfully, you can perform the minimization:
mdrun -s input_min.tpr -deffnm ubiquitin_min -v
By using the '-v' option, you can watch the minimization progress on the screen. Using the steepest descent method, small systems may equilibrate after only a few hundred steps; larger systems may may take several thousand steps. It mostly depends on the complexity of the system and the quality of the original starting structure. Once the minimization is complete (which should take around 500 steps), you will find that several files have been written to disk, all with the prefix 'ubiquitin_min'. They are as follows:
1.) ubiquitin_min.gro # system coordinates at the final step 2.) ubiquitin_min.trr # trajectory file 3.) ubiquitin_min.log # log file 4.) ubiquitin_min.edr # energy file
The first file, 'ubiquitin_min.gro', contains the fully minimized coordinates of the system (presuming the minimization was successful). The second file, 'ubiquitin_min.trr' contains a trajectory of the minimization. The frequency with which frames are written to the trajectory is specified in the .mdp file that was passed to grompp. You can view this in VMD by first loading the original coordinate file (protein_sol_nacl.gro), followed by the trajectory file. The log file contains information about the run parameters used for the minimization, as well as various system energies during the minimization. It is convenient to check the progress of long MD simulations by following the end of this file. Finally, ubiquitin_min.edr is a binary energy file that can be read by the GROMACS tool 'g_energy', yielding many pieces of valuable information. In this case, you can use it to check the potential energy of the system as a function of minimization step. Execute the following command:
g_energy -f ubiquitin_min.edr -o energy.xvg
A prompt will appear asking for a specific term you would like to analyze:
Select the terms you want from the following list by selecting either (part of) the name or the number or a combination. End your selection with an empty line or a zero. ------------------------------------------------------------------- 1 Bond 2 Angle 3 Proper-Dih. 4 Improper-Dih. 5 LJ-14 6 Coulomb-14 7 LJ-(SR) 8 Disper.-corr. 9 Coulomb-(SR) 10 Coul.-recip. 11 Potential 12 Pres.-DC 13 Pressure 14 Vir-XX 15 Vir-XY 16 Vir-XZ 17 Vir-YX 18 Vir-YY 19 Vir-YZ 20 Vir-ZX 21 Vir-ZY 22 Vir-ZZ 23 Pres-XX 24 Pres-XY 25 Pres-XZ 26 Pres-YX 27 Pres-YY 28 Pres-YZ 29 Pres-ZX 30 Pres-ZY 31 Pres-ZZ 32 #Surf*SurfTen 33 Mu-X 34 Mu-Y 35 Mu-Z 36 T-rest
Choose '11' for the potential energy, and hit return on an empty line to finish. A file called 'energy.xvg' will be written to file. This is a simple two-column data file that can be plotted in a program such as Grace:

Potential energy of system as a function of minimization step.
Convergence of the potential energy in a system to a large negative number (on the order of ~10^5) is a very good indication that the energy minimization was successful. If you find that for your system the potential energy has converged, and if the maximum force on any atom is below a reasonable tolerance cut-off, then it is okay to proceed to Equilibration MD. Otherwise, you may need to use other minimization methods (such as conjugate gradients or L-BFGS) to reach a lower energy well. The implementation of these are described in detail in the GROMACS manual.