Difference between revisions of "2010 DOCK tutorial with Streptavidin"
(→Enzyme Surface) |
(→Enzyme Surface) |
||
| Line 39: | Line 39: | ||
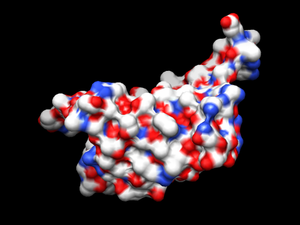
[[File:surface.png|thumb|Streptavidin enzyme surface]] | [[File:surface.png|thumb|Streptavidin enzyme surface]] | ||
| − | Recent versions of Chimera include a Write DMS tool that facilitates calculation of the molecular surface. Go to Tools -> Structure Editing -> Write DMS. Save the surface as 1DF8.rec.dms | + | Recent versions of Chimera include a Write DMS tool that facilitates calculation of the molecular surface. Go to Tools -> Structure Editing -> Write DMS. Save the surface as 1DF8.rec.dms. |
| − | + | The Write DMS algorithm will "roll" a small probe (default radius = 1.4 Angstroms) over the surface of the enzyme and calculate the surface normal at each point. Note that this can also be accomplished with a separate dms program, as described in DOCK tutorials from previous years. DMS (dot molecular surface) files are subsequently used as an input for sphgen. | |
===Spheres=== | ===Spheres=== | ||
Revision as of 10:35, 20 February 2010
Contents
About DOCK
DOCK was developed by Irwin D. "Tack" Kuntz, Jr., PhD and colleagues at UCSF. Please see the webpage at UCSF DOCK.
DOCK is a molecular docking program used in drug discovery. This program, given a protein active site and a small molecule, tries to predict the correct binding mode of the small molecule in the active site, and the associated binding energy. Small molecules with highly favorable binding energies could be new drug leads. This makes DOCK a valuable drug discovery tool. DOCK is typically used to screen massive libraries of millions of compounds against a protein to isolate potential drug leads. These leads are then further studied, and could eventually result in a new, marketable drug.
About Streptavidin & Biotin
Streptavidin is a tetrameric prokaryoke protein that binds the co-enzyme biotin with an extremely high affinity. The streptavidin monomer is composed of eight antiparallel beta-strands which folds to give a beta barrel tertiary structure. A biotin binding-site is located at one end of each β-barrel, which has a high affinity as well as a high avidity for biotin. Four identical streptavidin monomers associate to give streptavidin’s tetrameric quaternary structure. The biotin binding-site in each barrel consists of residues from the interior of the barrel, together with a conserved Trp120 from neighbouring subunit. In this way, each subunit contributes to the binding site on the neighboring subunit, and so the tetramer can also be considered a dimer of functional dimers.
Biotin is a water soluble B-vitamin complex which is composed of an ureido (tetrahydroimidizalone) ring fused with a tetrahydrothiophene ring. It is a co-enzyme that is required in the metabolism of fatty acids and leucine. It is also involved in gluconeogenisis.
Downloading the PDB complex (1DF8)
Download the neuraminidase file from here into your working directory. To get the monomer, download using the blue file arrow icon. Choose biological unit gz under download files folder at the left side. In that way you can downlad a biological gz file. If you download it by using "fetch by ID" in Chimera, you can only download a pdb gz file.
If the pdb gz file is downloaded the monomeric structure of neuraminidase with the bound ligand is seen.
However if the biological gz file is downloaded a tetramer complex of neuraminidase is seen.
When docking, we will only choose a monomer of the whole protein and its ligands instead of the whole tetramer. And it will be painful if you follow the procedure below, use vi to prepare the enzyme and ligand. Because you need to make sure the ligand you extract from the whole pdb files is right the one that match the monomer you've chosen.
Preparing the Enzyme and Ligand in Chimera
To delete the water molecules, etc. go to tools->struc. edit--> dock prep. Check all boxes and click ok. Save as 1DF8.dockprep.mol2 To separate the ligand: select--->residue-->BTN. Then go to actions-->atoms--->delete Save file: 1DF8.receptor.mol2 Select--->chemistry-->element-->H and delete. Save as PDB file 1DF8.receptor.noH.pdb close and open dockprep file again. select protein and delete. Now have ligand only. Save as 1DF8.lig.mol2
Generation of Enzyme Surface, Spheres, and Grid for DOCK
Enzyme Surface
To generate an enzyme surface, first open the receptor pdb file where the hydrogen atoms were removed (1DF8.rec.noH.pdb). Next, go to Actions -> Surface -> Show.
Recent versions of Chimera include a Write DMS tool that facilitates calculation of the molecular surface. Go to Tools -> Structure Editing -> Write DMS. Save the surface as 1DF8.rec.dms.
The Write DMS algorithm will "roll" a small probe (default radius = 1.4 Angstroms) over the surface of the enzyme and calculate the surface normal at each point. Note that this can also be accomplished with a separate dms program, as described in DOCK tutorials from previous years. DMS (dot molecular surface) files are subsequently used as an input for sphgen.
Spheres
To generate spheres file, we need to use command line program called sphgen. To run the sphgen, we need a input file named INSPH. The content of INSPH is like this
1DF8_receptor.dms R X 0.0 4.0 1.4 1DF8_receptor.sph
1DF8_receptor.ms is the surface file we got from the previous step. (Change the file name to what you named it before). 1DF8_receptor.sph is the spheres file we want to generate in this step.
Use this command to generate spheres file
sphgen -i INSPH -o OUTSPH
You should get the OUTSPH similar to this
density type = X reading 1DF8_receptor.dms type R # of atoms = 881 # of surf pts = 10771 finding spheres for 1DF8_receptor.dms dotlim = 0.000 radmax = 4.000 Minimum radius of acceptable spheres? 1.4000000 output to 1DF8_receptor.sph clustering is complete 27 clusters
You can open the spheres file (1DF8_receptor.sph). There are over 700 atoms in this file. However, we're only interested in docking the ligand into the active site. To get better efficiency, we can select those sphere atoms in the active site by using sphere_selector command.
sphere_selector 1DF8_receptor.sph 1DF8_ligand.mol2 10.0
Where 1DF8_ligand is the ligand file. 10.0 is the distance cutoff. You should get a file called selected_spheres.sph. You can open it. The number of atoms is drop down to 47.