Difference between revisions of "2013 DOCK tutorial with Orotodine Monophosphate Decarboxylase"
Stonybrook (talk | contribs) (→V. Docking a Single Molecule for Pose Reproduction) |
Stonybrook (talk | contribs) (→IV. Generating Box and Grid) |
||
| Line 156: | Line 156: | ||
==IV. Generating Box and Grid== | ==IV. Generating Box and Grid== | ||
| + | '''BOX''' | ||
| + | |||
| + | Create a box around the active site, which set the out boundaries of energy calculation. | ||
| + | |||
| + | First create a directory where you put all your grid and box files. | ||
mkdir 03.box.grid | mkdir 03.box.grid | ||
cd 03.box.grid | cd 03.box.grid | ||
| + | |||
| + | Then you need to create a "showbox.in" file to generate the box. | ||
| + | |||
| + | $touch showbox.in | ||
| + | |||
| + | Edit this file as following: | ||
| + | |||
| + | Y | ||
| + | 8.0 | ||
| + | ../02.surface-sphgen/selected_spheres.sph | ||
| + | 1 | ||
| + | 1LOQ.box.pdb | ||
| + | |||
| + | To feed the answers to the questions, run | ||
| + | $showbox < showbox.in | ||
| + | |||
| + | [[File:boxexample.png|frame|Box Image]] | ||
| + | |||
| + | |||
| + | '''GRID''' | ||
| + | |||
| + | Calculate electrostatic and VDW potential of the receptor at each grid point. Method to speed up evaluation of the energy scoring function. | ||
| + | |||
| + | Use grid program to generate the grid and answer the questions as following: | ||
| + | |||
| + | compute_grids yes | ||
| + | grid_spacing 0.4 | ||
| + | output_molecule no | ||
| + | contact_score no | ||
| + | energy_score yes | ||
| + | energy_cutoff_distance 9999 | ||
| + | atom_model a | ||
| + | attractive_exponent 6 | ||
| + | repulsive_exponent 9 | ||
| + | distance_dielectric yes | ||
| + | dielectric_factor 4 | ||
| + | bump_filter yes | ||
| + | bump_overlap 0.75 | ||
| + | receptor_file ../01.dockprep/1LOQ.receptor.mol2 | ||
| + | box_file ../03.box.grid/1LOQ.box.pdb | ||
| + | vdw_definition_file ../00.files/vdw_AMBER_parm99.defn | ||
| + | score_grid_prefix 1LOQ.grid | ||
| + | |||
| + | [[File:gridexample.png|frame|Box Image]] | ||
==V. Docking a Single Molecule for Pose Reproduction== | ==V. Docking a Single Molecule for Pose Reproduction== | ||
Revision as of 22:06, 5 March 2013
For additional Rizzo Lab tutorials see DOCK Tutorials. Use this link Wiki Formatting as a reference for editing the wiki. This tutorial was developed collaboratively by the AMS 536 class of 2013, using DOCK v6.6.
Contents
- 1 I. Introduction
- 2 II. Preparing the Receptor and Ligand
- 3 III. Generating Receptor Surface and Spheres
- 4 IV. Generating Box and Grid
- 5 V. Docking a Single Molecule for Pose Reproduction
- 6 VI. Virtual Screening
- 7 VII. Running DOCK in Serial and in Parallel on Seawulf
- 8 VIII. Frequently Encountered Problems
I. Introduction
DOCK
DOCK is a molecular docking program used in drug discovery. It was developed by Irwin D. Kuntz, Jr. and colleagues at UCSF (see UCSF DOCK). This program, given a protein binding site and a small molecule, tries to predict the correct binding mode of the small molecule in the binding site, and the associated binding energy. Small molecules with highly favorable binding energies could be new drug leads. This makes DOCK a valuable drug discovery tool. DOCK is typically used to screen massive libraries of millions of compounds against a protein to isolate potential drug leads. These leads are then further studied, and could eventually result in a new, marketable drug. DOCK works well as a screening procedure for generating leads, but is not currently as useful for optimization of those leads.
DOCK 6 uses an incremental construction algorithm called anchor and grow. It is described by a three-step process:
- Rigid portion of ligand (anchor) is docked by geometric methods.
- Non-rigid segments added in layers; energy minimized.
- The resulting configurations are 'pruned' and energy re-minimized, yielding the docked configurations.
Orotodine Monophosphate Decarboxylase
The protein receptor which is the subject of this tutorial is orotodine monophosphate decarboxylase (OMP), a homodimeric protein from the organism Methanobacterium thermoautotrophicum. OMP is involved in the biosynthesis of several pyrimidines including uridine monophosphate (UMP), the ligand used in this tutorial. UMP is a pyrimidine-based nucleotide monomer of RNA. The structure used for this tutorial can be found at the Protein Data Bank under accenssion number 1LOQ.
Organizing Directories
While performing docking, it is convenient to adopt a standard directory structure / naming scheme, so that files are easy to find / identify. For this tutorial, we will use something similar to the following:
~username/AMS536/dock-tutorial/00.files/
/01.dockprep/
/02.surface-sphgen/
/03.box-grid/
/04.dock/
/05.mini-virtual-screen/
/06.database-filter/
/07.virtual-screen/
In addition, most of the important files that are derived from the original crystal structure will be given a prefix that is the same as the PDB code, '1LOQ'. The following sections in this tutorial will adhere to this directory structure / naming scheme.
II. Preparing the Receptor and Ligand
(Ye and Weiliang)
PDB file downloading (1LOQ): go to PDB homepage(http://www.rcsb.org/pdb/home/home.do ) enter the protein ID (1LOQ) in the PDB, click Download Files in the top-right of the webpage, then select PDB File (text). In the new window, save the file in Downloads.
Generating files for ligand and receptor:
In this section, we will create four new files in the 00-original-files folder:
1LOQ.dockprep.mol2 - ligand molecule, with hydrogens and am 1-bcc partial charges.
1LOQ.receptor.mol2 - receptor molecule, with hydrogens and amber charges.
1LOQ.receptor.noH.mol2 - receptor without hydrogen atoms.
1LOQ.ligand.mol2 - ligand only.
Firstly please copy the pdb file in 00-original-files folder;then open the pdb file in the promt "vim 1LOQ.pdb";in the pdb file, change all residues "U" by "LIG" starting at 2082 line; here is the comment for you to search in the vim sheet to change the ligand U into LIG to ensure the ligand can be read by Chimera:
%s/ U/LIG/gc
g is short for global check and c is short for checking before having changes
The preparation will be shown in Chimera. Open Chimera by typing Chimera at the prompt if you are on "herbie"; Click Open in File menu and find the file "1LOQ.pdb";
To delete water molecules/other ligands, click Structure Editing in Tools manu and click Dock Prep. Check all boxes and click okey to the end.
Or go to Select->residue->HOH, then go to actions->delete then u remove water in the original molecule.
then add H to the molecule:tools->Structure editing->add H
Next, to add charge to the ligand, go to Select->residue->LIG,then go to tools->Structure editing->->add charge, and then chose "AMBER ff99SB" in charge model -> select AM1-BCC as charge method, then assign residue LIG net charge for -1
To create a receptor file:
Open 1LOQ.dockprep.mol2, click Select -> Residue -> LIG. Then click Actions -> Atoms -> Delete. Save the file as 1LOQ.receptor.mol2.
To create a receptor file with no hydrogen atoms:
Open 1LOQ.dockprep.mol2, click Select -> Chemistry -> Element -> H -> Delete. Save the file as 1LOQ.receptor.noH.mol2.
To create a Ligand file:
Open 1LOQ.dockprep.mol2, click Select -> Chain -> Delete. Save the file as 1LOQ.ligand.mol2.
III. Generating Receptor Surface and Spheres
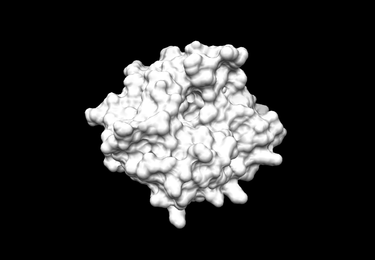
Receptor Surface
In order to generate the receptor surface one can perform the following steps (working in Chimera):
- Go File -> Open and choose the (generated at previous step) pdb file of the protein containing no Hydrogens (1LOQ.receptor.noH.pdb)
- Further, Actions -> Surface -> Show
- Go Tools -> Structure Editing -> Write DMS in order to obtain a dms file, which we will need while placing spheres
- In the window appeared save the surface as 1LOQ.receptor.dms
Placing Spheres
In order to place the spheres on the receptero surface, the following steps are required:
1. Create a file INSPH (an input file we will need further) and fill it out as follows (description of parameters to the right), then save it.
1LOQ.receptor.dms #molecular surface file that we got from previous step R #sphere outside of surface (R) or inside surface (L) X #specifies subset of surface points to be used (X=all points) 0.0 #prevents generation of large spheres with close surface contacts(defalut=0.0) 4.0 #maximum sphere radius in angstroms (default=4.0) 1.4 #minimum sphere radius in angstroms (default=radius of probe) 1LOQ.receptor.sph #clustered spheres file that we want to generate
Make sure that your file doesn't contain any blank lines or the comments (denoted by # ) from above, otherwise it won't work.
2. Run the sphgen program from the command line:
sphgen -i INSPH -o OUTSPH
OUTSPH is the output file containing the information on the sphgen run, while 1LOQ.receptor.sph is the file where the information about spheres generated could be found.
3. (optional) If one wants to have a look at the spheres received as an outcome of sphgen, one should run showsphere, by typing in the commandline:
showsphere
Later on the following questions will be asked:
Enter name of sphere cluster file: Enter cluster number to process (<0 = all): Generate surfaces as well as pdb files (<N>/Y)? Enter name for output file prefix: Process cluster 0 (contains ALL spheres) (<N>/Y)?
As the questions have been answered the program will generate the spheres, writing the data in a pdb file, afterwards the spheres may be observed by opening this file in Chimera.
4. Run sphere_selector program form the commandline as follows:
sphere_selector 1LOQ.receptor.sph ../01-dockprep/1LOQ.ligand.mol2 8.0
This program selects the spheres within a user-defined radius (8.0 here) of a target molecule from the previously obtained file 1LOQ.receptor.sph. As a result a new file selected_spheres.sph will be generated. Again, the spheres might be visualized by using showsphere and Chimera, but formally, we prepared the data for moving further.
5. (optional) In order to see the spheres:
- Run showsphere, answering the questions as follows:
selected_spheres.sph -1 N output_spheres_selected N
- Launch Chimera, go File -> Open, choose 1LOQ.receptor.noH.pdb
- Go File -> Open, choosing output_spheres_selected.pdb
- Go Select -> Residue -> SPH
- Finally, Actions -> Atoms/Bonds -> sphere
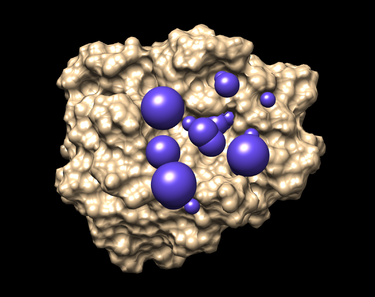
The picture like one placed below will be obtained afterwards.
IV. Generating Box and Grid
BOX
Create a box around the active site, which set the out boundaries of energy calculation.
First create a directory where you put all your grid and box files.
mkdir 03.box.grid cd 03.box.grid
Then you need to create a "showbox.in" file to generate the box.
$touch showbox.in
Edit this file as following:
Y 8.0 ../02.surface-sphgen/selected_spheres.sph 1 1LOQ.box.pdb
To feed the answers to the questions, run
$showbox < showbox.in
GRID
Calculate electrostatic and VDW potential of the receptor at each grid point. Method to speed up evaluation of the energy scoring function.
Use grid program to generate the grid and answer the questions as following:
compute_grids yes grid_spacing 0.4 output_molecule no contact_score no energy_score yes energy_cutoff_distance 9999 atom_model a attractive_exponent 6 repulsive_exponent 9 distance_dielectric yes dielectric_factor 4 bump_filter yes bump_overlap 0.75 receptor_file ../01.dockprep/1LOQ.receptor.mol2 box_file ../03.box.grid/1LOQ.box.pdb vdw_definition_file ../00.files/vdw_AMBER_parm99.defn score_grid_prefix 1LOQ.grid
V. Docking a Single Molecule for Pose Reproduction
Create the input file for docking:
touch dock.in
Execute Dock6:
dock6 -i dock.in
This will generate the questions to set the parameters. As an example, for the rigid body docking the parameters will be as follows:
ligand_atom_file ../01.dockprep/1LOQ.ligand.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd no use_database_filter no orient_ligand no use_internal_energy no flexible_ligand no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary yes grid_score_secondary no grid_score_rep_rad_scale 1 grid_score_vdw_scale 1 grid_score_es_scale 1 grid_score_grid_prefix ../03.box-grid/1LOQ.grid multigrid_score_secondary no dock3.5_score_secondary no continuous_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_descriptor_score_secondary no amber_score_secondary no minimize_ligand no atom_model all vdw_defn_file ../00.files/vdw_AMBER_parm99.defn flex_defn_file ../00.files/flex.defn flex_drive_file ../00.files/flex_drive.tbl ligand_outfile_prefix output write_orientations no num_scored_conformers 1 rank_ligands no
The rigid docking will result in a single-ligand pose. Open Chimera to visualize the docking result. After opening the receptor file in surface view, open the ligand file (named "output_scored.mol2") by going under "Tools--> Surface/Binding Analysis--> ViewDock".
For flexible docking, change input parameters. These changes will generate additional questions. Some changes to parameters are shown below:
calculate_rmsd yes orient_ligand yes flexible_ligand yes use_internal_energy yes minimize_ligand yes num_scored_conformers 10
This flexible ligand protocol can be used with multiple ligand input files. The docked ligand poses can be compared using RMSD (Root mean squared Deviation). RMSD is a measurement of the distance between atoms on the original crystal structure ligand with the newly generated ligand conformation. An RMSD under 2 Angstroms would indicate a good conformation. Below are two examples of a final docked ligand in the receptor active site pocket. The worst and best energy scored ligands are shown below. The original ligand is in cyan and the docked ligand is colored in heteroatom.
VI. Virtual Screening
Virtual Screening Introduction
A virtual screen of various ligand allows for the comparison of both qualitative (e.g. position in binding site) and quantitative (e.g. energy scores) data pertaining to the each screened ligand with an originally docked molecule. Virtual screening is often used as a method to cut the cost of experimentation by narrowing down the ligands within a database and predicting which will exhibit the most favorable binding to a specific protein (with a pre-determined .grid file).
Creating a Filter for the Virtual Screen
When first beginning the virtual screen, a filter should be specified in order save computational power by only screening potentially effective ligands within the directory 06.database-filter/. This involves limiting several variables of ligands specified in a .mol2 file. To limit the ligands from the original database, a file dock_filter.in is created. Here are the contents of the file:
ligand_atom_file /home/wjallen/AMS536/multi-mol2-files/cdiv_p0.0.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd no use_database_filter yes dbfilter_max_heavy_atoms 50 dbfilter_min_heavy_atoms 0 dbfilter_max_rot_bonds 10 dbfilter_min_rot_bonds 0 dbfilter_max_molwt 9999.0 dbfilter_min_molwt 0.0 dbfilter_max_formal_charge -1.0 dbfilter_min_formal_charge -2.0 orient_ligand no use_internal_energy no flexible_ligand no bump_filter no score_molecules no atom_model all vdw_defn_file ../00.files/vdw_AMBER_parm99.defn flex_defn_file ../00.files/flex.defn flex_drive_file ../00.files/flex_drive.tbl ligand_outfile_prefix filtered write_orientations no num_scored_conformers 1 rank_ligands no
In this file some important characteristics of the ligands that are limited are dbfilter_max_heavy_atoms, dbfilter_max_rot_bonds, dbfilter_max_molwt, and both dbfilter_max_formal_charge and dbfilter_min_formal_charge. As show in the text of the file, a file called filtered_scored.mol2 will be created with all of the ligands from the original database that fit into the specified limits of number of heavy atoms, number of rotatable bonds, molecular weight, and formal charge.
Proceeding to the Virtual Screen
After creating a new directory, 07.virtual-screen/, a dock.in file must be created. So as not to confuse the virtual screen file with the original dock.in of the docked ligand, the new file created is called dock_vs.in. Running this file in dock6 allows us to fill in the important details of the file that terminate in a virtual screening of the input ligand.
[Note to self: add text from dock_vs.in]
VII. Running DOCK in Serial and in Parallel on Seawulf
The Seawulf Cluster is a 470-processor Linux Cluster capable of highly parallel processing. This parallel processing allows dock virtual screens to be completed in a fraction of the time as a single processor.
If you are docking multiple ligands, you can use more than one processor in parallel mode, but you should never use more processors than you have ligands. Before we can run DOCK on Seawulf, we need to copy the proper files from Herbie to Seawulf. If we CD into the AMS536 folder we can use the following command from the mathlab computer to copy all of the dock-tutorial files
$scp -r /dock-tutorial/ username@herbie.mathlab.sunysb.edu:~/AMS536/
Now we have all of our DOCK preparation files and folders on the seawulf cluster.
Running DOCK in Serial on a Single Processor
Running on a single processor is very similar to running dock on the mathlab comptuer.
If you make a file called qsub.csh with the text:
#!/bin/tcsh #PBS -l nodes=1:ppn=1 #PBS -l walltime=01:00:00 #PBS -N dock6 #PBS -M user@ic.sunysb.edu #PBS -j oe #PBS -o pbs.out cd /nfs/user03/zfoda/AMS536/dock-tutorial/07.virtscreen /nfs/user03/wjallen/local/dock6/bin/dock6 -i dock.in -o dock.out
An explanation of the commands:
#!/bin/tcsh #Execute script with tcsh #PBS -l nodes=1:ppn=1 #Use one node, and one processor per node, so one single processor #PBS -l walltime=01:00:00 #Allow 1 hour for your job run #PBS -N dock6 #Name of your job #PBS -M user@ic.sunysb.edu #Get an email notifying you when your job is completed #PBS -j oe #Combine the output and error streams into a single output file #PBS -o pbs.out #Name of your output file cd /nfs/user03/zfoda/AMS536/dock-tutorial/07.virtscreen #Change to your home directory and folder with dock files /nfs/user03/wjallen/local/dock6/bin/dock6 -i dock.in -o dock.out #Specifies path to dock executable and provide input and output filenames
To submit the experiment use the command:
qsub qsub.csh
You will have submitted a DOCK experiment to the seawulf queue.
See also PBS commands.
Running DOCK in Parallel using MPI
In order to run DOCK in parallel you have to use a slightly different build of DOCK6 called dock6.mpi. Message passing interface (MPI) is basically a program that allows programs like DOCK to run in parallel.
So, make another file called qsub.vs.csh with the contents:
#!/bin/tcsh #PBS -l nodes=4:ppn=2 #PBS -l walltime=24:00:00 #PBS -N screen #PBS -o qsub.log #PBS -j oe #PBS -V cd /nfs/user03/username/AMS536/dock-tutorial/07.virtscreen mpirun -np 8 /nfs/user03/wjallen/local/dock6/bin/dock6.mpi -i dockvs.in -o dockvs.out
As you can see there are two major changes:
#PBS -l nodes=4:ppn=2 #Use 4 nodes, and 2 processors per node, so 8 processors
Note: since one processor is used to distribute the processes, this will run DOCK as 7 (n-1) parallel processes.
mpirun -np 8 /nfs/user03/wjallen/local/dock6/bin/dock6.mpi -i dockvs.in -o dockvs.out #this line uses mpi to run dock.mpi on multiple processors
And then we can run:
qsub qsub.vs.csh