Difference between revisions of "2023 DOCK tutorial 3 with PDBID 2P16"
Stonybrook (talk | contribs) (→Introduction) |
Stonybrook (talk | contribs) (→Introduction) |
||
| Line 11: | Line 11: | ||
* Cartesian Minimization | * Cartesian Minimization | ||
| − | For this tutorial, please ensure that you | + | For this tutorial, please ensure that you have DOCK6.10 installed and accessible in your path, which you can check using the commands: |
| − | <code> | + | <code>which dock</code> |
| − | which dock | + | <code>echo $PATH</code> |
| − | echo $PATH | ||
| − | </code> | ||
=== '''2P16 System''' === | === '''2P16 System''' === | ||
| Line 31: | Line 29: | ||
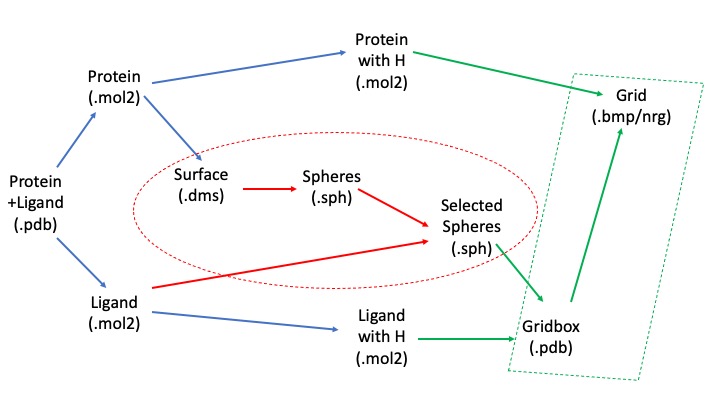
[[File:docking_flowchart.jpg]] | [[File:docking_flowchart.jpg]] | ||
| + | |||
| + | We will begin with the pdb file, clean up and separate the protein and ligand. We will use the unprotonated forms to calculate spheres, which, along with our protonated form, we will use to calculate the scoring grid. The blue arrows represent transformations we can make suing Chimera, the red ones using SPHGEN and sphere_selector, and the green ones using showbox and grid commands. | ||
== '''Protein and Ligand Preparation''' == | == '''Protein and Ligand Preparation''' == | ||
Revision as of 23:22, 23 March 2023
Contents
Introduction
Docking is a molecular modeling program that useful for drug discovery. Broadly speaking, it computes optimal binding conformations and affinities of a small molecule to its target protein. The discovery of new low-energy ligand binding modes to the active site of the protein can help discover new or optimize existing drugs.
DOCK6
DOCK6 was initially developed at UCSF as an improvement over traditional rigid docking methods and now is maintained by several groups across 3 universities. In this tutorial we will be exploring the following functions:
- Docking- rigid, flexible, and fixed anchor
- Virtual Screening
- Cartesian Minimization
For this tutorial, please ensure that you have DOCK6.10 installed and accessible in your path, which you can check using the commands:
which dock
echo $PATH
2P16 System
2p16 refers to the pdb code for the crystal structure of Factor Xa in complex with its inhibitor Apixaban.
Factor Xa plays a key role in the formation of blood clots and is therefore a drug target for anticoagulants.
The drug Apixaban is a second-generation therapy developed by Bristol Myers Squibb- it has fewer drug interactions than the more traditional Heparin and Warfarin, and therefore can be used in patients with additional complications. A docking study of this system may elucide the mechanism of this increased specificity and help design the next generation of specific inhibitors.
Directory Structure
In order to understand the necessary directories it's important to understand the workflow of this project:
We will begin with the pdb file, clean up and separate the protein and ligand. We will use the unprotonated forms to calculate spheres, which, along with our protonated form, we will use to calculate the scoring grid. The blue arrows represent transformations we can make suing Chimera, the red ones using SPHGEN and sphere_selector, and the green ones using showbox and grid commands.
Protein and Ligand Preparation
Protein Preparation
Ligand Preparation
Surface Spheres Generation
Grid Box
Grid Generation
Box Generation
Ligand Minimization
Footprint Analysis
DOCKING
The goal of docking in this tutorial is to reproduce the crystal structure from x-ray diffraction. This is called “pose reproduction”; this can be used to validate the docking software and parameters of use within the software. If the pose reproduction is not successful, it might be possible that: 1.the parameters to dock the ligand needs to be changed, 2.preparation of ligand or receptor went wrong, 3.the system itself is not suitable to reproduce in the software. Having a preliminary docking pose before getting into virtual screening–a procedure docking thousands of molecules–would be helpful to validate the results you will get in the future. In DOCK6 (at the point of 2020), the docking is successful if the RMSD of the docked pose is within 2.0Å from the crystal pose. In this section, we will be working on (i)rigid docking, (ii)flexible docking, (iii)fixed anchor docking.
Rigid Docking
Rigid docking does not conduct pose searching with dihedral rotations or bond length perturbations. It takes 3 translational and 3 rotational degrees of freedom to do its pose searching. Therefore, rigid docking is less computationally expensive compared to flex docking. Since we already have the crystal structure of the ligand bound at the active site, it is expected to have a good reproduction using rigid docking.
The first step is to create an input file by rigid.in
Insert the following into that file:
conformer_search_type rigid use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100 ligand_atom_file 2p16.ligand.min_scored.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd yes use_rmsd_reference_mol yes rmsd_reference_filename 2p16.ligand.min_scored.mol2 use_database_filter no orient_ligand yes automated_matching yes receptor_site_file selected_spheres.sph max_orientations 2000 critical_points no chemical_matching no use_ligand_spheres no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary yes grid_score_secondary no grid_score_rep_rad_scale 1 grid_score_vdw_scale 1 grid_score_es_scale 1 grid_score_grid_prefix grid multigrid_score_secondary no dock3.5_score_secondary no continuous_score_secondary no footprint_similarity_score_secondary no pharmacophore_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand yes simplex_max_iterations 1000 simplex_tors_premin_iterations 0 simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1.0 simplex_trans_step 1.0 simplex_rot_step 0.1 simplex_tors_step 10.0 simplex_random_seed 0 simplex_restraint_min no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl ligand_outfile_prefix rigid.out write_orientations no num_scored_conformers 1 rank_ligands no
Make sure to (i)have all the files you need (selected_spheres.sph and 2p16.ligand.min_scored.mol2) and (ii)inputs are correct.
Run the input file with the following command:
dock6 -i rigid.in -o rigid.out
After the job is complete, you should see a file named “rigid.out_scored.mol2”. This is the output that shows the docked poses (if you rank it, it will show from the highest to lowest docking score). If you open the file, in the first part, you will see general information about the docked pose including the grid scores. Make sure that the output file is not empty.
The pose can be visualized using Chimera.
After you load the protein you prepared (simply open the file), you can go to:
Tools -> Surface/Binding Analysis -> ViewDock -> *choose the file* -> Dock4,5, or 6.
Along with the visualized pose, you will see a small window popping out with the quantitative information as you saw in the first part of the output file.
Since we took a crystal structure of the ligand, minimized, and rigidly docked into the same protein, it is expected that the produced pose is well-reproduced.
If you want to check the reproducibility, you could check out the RMSD (root-mean-square-deviation) values for the pose.
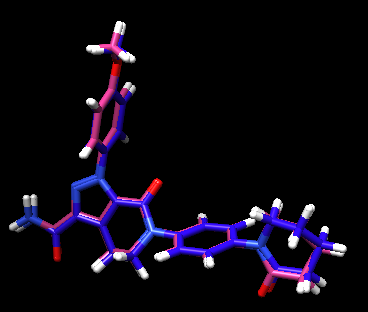
The crystal reference ligand is colored in pink
From the result pose, you can see that the pose reproduction was not successful. From the scored.mol2 file, the RMSD was indicated to be 7.4 Å, which is above 2.0 Å cutoff for a successful pose reproduction. Although this was an unexpected results from the initial assumption, but we can try using two other methods: flex and fixed anchor docking.
Fixed Anchor Docking
As it is said “fixed anchor” docking, this is a docking method that docks while maintaining the orientation of the initially placed structure. For this reason, “comformer_search_type” is flexible, while “write orientation” is turned off. This allows more flexibility in its conformational change compared to rigid docking methods while maintaining the general placement. You can also pick specific fragments within the molecules to be the anchor.
Not to mess up the outputs with other types of docking methods, make a new folder by
mkdir anchor_dock
In this directory, create an input file:
vi fixed.in
Insert the following:
conformer_search_type flex write_fragment_libraries no user_specified_anchor no limit_max_anchors no min_anchor_size 5 pruning_use_clustering yes pruning_max_orients 10000 pruning_clustering_cutoff 100 pruning_conformer_score_cutoff 100.0 pruning_conformer_score_scaling_factor 1.0 use_clash_overlap no write_growth_tree no use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 ligand_atom_file 2p16.charge.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd yes use_rmsd_reference_mol yes rmsd_reference_filename 2p16.charge.mol2 use_database_filter no orient_ligand no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary yes grid_score_secondary no grid_score_rep_rad_scale 1 grid_score_vdw_scale 1 grid_score_es_scale 1 grid_score_grid_prefix grid multigrid_score_secondary no dock3.5_score_secondary no continuous_score_secondary no footprint_similarity_score_secondary no pharmacophore_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand yes minimize_anchor yes minimize_flexible_growth yes use_advanced_simplex_parameters no simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1 simplex_trans_step 1 simplex_rot_step 0.1 simplex_tors_step 10 simplex_anchor_max_iterations 500 simplex_grow_max_iterations 500 simplex_grow_tors_premin_iterations 0 simplex_random_seed 0 simplex_restraint_min no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl ligand_outfile_prefix 2P16_fixed write_orientations no num_scored_conformers 1 rank_ligands no
Start the docking by:
dock6 -i fixed.in -o fixed.out
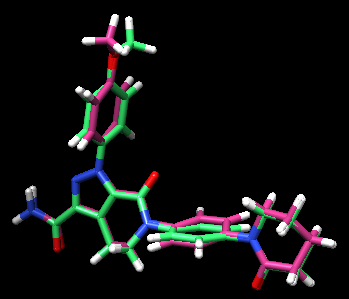
The resulting docked pose:
The RMSD in this result was 0.22 Å, which is well within 2.0 Å (success cutoff). From the rigid docking result, we could see that the orientation of the entire molecule was flipped from the minimized crystal structure. By having a part of the molecule anchored, it seemed that the big rotation of this molecule did not occur. For a case of new compounds, ones that do not have a crystal structures, this method allows more chemical exploration.
Flex Docking
Finally, flex docking is a docking method that allows flexibility in the rotatable bonds in the ligand and orientation of the compound. This method has the largest exploration of the pose for the ligand, although the calculation time takes the longest. To speed up the calculation time, it is necessary to use cpus provided in seawulf. To use the cpus, we need to submit our run to the queue.
Create a directory:
mkdir flex_dock
First, start with creating an input file, flex.in.
conformer_search_type flex write_fragment_libraries no user_specified_anchor no limit_max_anchors no min_anchor_size 5 pruning_use_clustering yes pruning_max_orients 5000 pruning_clustering_cutoff 2500 pruning_conformer_score_cutoff 100.0 pruning_conformer_score_scaling_factor 1.0 use_clash_overlap no write_growth_tree no use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 ligand_atom_file ../Rigid_dock/2p16.ligand.min_scored.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd yes use_rmsd_reference_mol yes rmsd_reference_filename ../Rigid_dock/2p16.ligand.min_scored.mol2 use_database_filter no orient_ligand yes automated_matching yes receptor_site_file ../Rigid_dock/selected_spheres.sph max_orientations 1000 critical_points no chemical_matching no use_ligand_spheres no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary yes grid_score_secondary no grid_score_rep_rad_scale 1 grid_score_vdw_scale 1 grid_score_es_scale 1 grid_score_grid_prefix ../Rigid_dock/grid multigrid_score_secondary no dock3.5_score_secondary no continuous_score_secondary no footprint_similarity_score_secondary no pharmacophore_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand yes minimize_anchor yes minimize_flexible_growth yes use_advanced_simplex_parameters no simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1.0 simplex_trans_step 1.0 simplex_rot_step 0.1 simplex_tors_step 10.0 simplex_anchor_max_iterations 500 simplex_grow_max_iterations 500 simplex_grow_tors_premin_iterations 0 simplex_random_seed 0 simplex_restraint_min no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl ligand_outfile_prefix flex.out write_orientations yes num_scored_conformers 10 write_conformations yes cluster_conformations yes cluster_rmsd_threshold 2.0 rank_ligands no
Now that we have our .in file, we need to make a slurm script to run it. To make a script, first run the following:
vi flex.sh
Inside of this file, copy-paste the following:
#!/bin/bash #SBATCH --time=10:00:00 #SBATCH --nodes=1 #SBATCH --ntasks=28 #SBATCH --job-name=2p16_flex_dock #SBATCH --output=2p16_flex_dock #SBATCH -p long-28core
dock6 -i flex.in -o 2p16_flex.out
You can submit the queue by
sbatch flex.sh
Once complete, you will see three .mol2 files: (1)flex.out_scored.mol2: best poses scored from the highest to lowest (2)flex.out_conformers.mol2: poses with all conformations of the ligand (3)flex.out_orients.mol2: poses with all orientations of the ligand
To check the best pose, check the flex.out_scored.mol2. You can open it on Chimera using viewdock. Compare with the crystal structure - is the pose with freedom of orientation and rotatable bonds matching with the binding pose in real life?
By looking at the comparison (green is the flex output), you can see that the pose reproduction went successfully. The output file showed the RMSD of 0.92 Å, which also indicates a success. This structure was found in the first in scored.mol2 showing that this pose was in fact the best pose after rotational bond and orientation searching.