2023 DOCK tutorial 1 with PDBID 4S0V
Contents
Introduction
This tutorial will walk you through the steps necessary for using the DOCK software package. Many drugs are small molecular compounds that attach, or bind, to a protein in our bodies to change how that protein functions. By changing the function of a protein we can treat disease and help people manage symptoms of disorders. Traditionally drug discovery was done through a type of "trial and error" process called High Throughput Screening. Scientists would chemically make, or buy, thousands of small compounds and expose them to cells. They would then observe how the cells responded, either favorably/unfavorably/no effect. This method is time consuming and expensive. It would be better if the scientific community could "virtually screen" these molecules using a computer before creating/buying them - thereby focusing the cost and effort on those which showed the most promising computational results. The DOCK software brings this drug discovery process into the 21st century and uses computers to bind these small molecular compounds to a protein and evaluate the results. DOCK uses algorithms to bring together the small molecule, known as the ligand, and the larger protein, and "DOCK" them together. Our tutorial will walk you through preparing a protein and ligand for docking using an example complex from the protein data bank (https://www.rcsb.org/), complex # 4S0V.
The following steps will be followed:
- Setting up your environmnet
- Downloading a protein from the PDB database
- Determining if there are any missing loops and if they need to be modeled
- Preparing the ligand
- Preparing the protein
Learning Objectives
- Understand why DOCK was created and its current role in drug design
- Gain the ability perform virtual screening of small molecular compounds to a protein from the Protein Data Base (https://www.rcsb.org/)
Setting Up Your Environment
TK
Downloading a protein from the PDB database
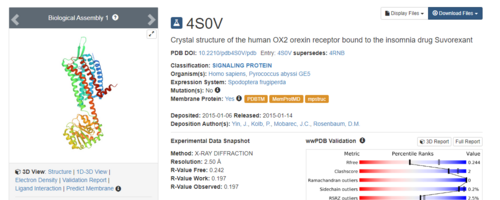
To begin we need to download protein complex #4S0V from the PDB. The right side of the top banner has a search bar:
.Simply type 4S0V into the search bar and the protein complex will be displayed.
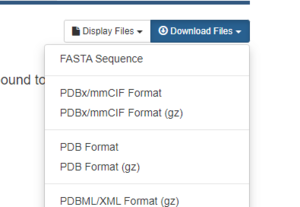
On the right hand side, click on Download Files, then PDB Format.
That's it! The pdb file that we will be working with is now downloaded onto your local computer.
Preparation of the ligand and protein
The following steps will show you how to prepare the protein and ligand structures to be used with DOCK. All steps in this section will be done using Chimera. If you are unfamiliar with Chimaera, please see our tutorials here. These steps are very important - if your initial structure is not prepared properly, all downstream analysis can potentially be incorrect. This section will show you how to:
- Evaluate the structure to determine if there are any missing loops/water molecules to be removed/ions that need to be removed
- How to prepare the protein structure
- How to prepare the ligand structure
Evaluating the Structure
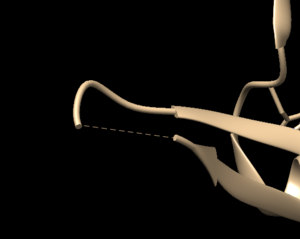
Open the previously downloaded .pdb file into Chimera. The first thing you want to look for are missing loops. A missing loop will be indicated by a dashed line in the structure:
The first decision you'll need to make is if these missing loops are important in your model or not. This decision is made by determining if the missing loop is close to the binding site. If it is far enough away, it probably won't affect the dynamics of the protein/ligand interaction and can be left alone. If the missing section is close to the binding site, you may want to fix it to more accurately model the binding site and the protein/ligand interaction.
To determine the distance between the missing loop and the binding site:
- Select an atom at the section of missing loop
- Select another atom near the binding site
- Go to Tools → Structure Analysis → Distances
To model a missing loop choose Tools → Surface Editing → Model/Refine Loops