2021 Denovo tutorial 3 with PDBID 1S19
This tutorial continues where we left off from the virtual screening of 1S19, a structure of the vitamin D nuclear receptor bound to the side chain analogue calcipotriol.
Contents
Focused Denovo Design with PDB 1S19
In this section, we will generate new ligand structures using a fragment library. It will require the use of some of the output files from the Virtual Screen tutorial (found here).
Fragment Libraries
We will use a focused fragment library file based on the original ligand. To begin, we will create a new directory for this.
mkdir fraglib
Create a new file called fragment.in with the following inputs.
conformer_search_type flex write_fragment_libraries yes fragment_library_prefix fraglib fragment_library_freq_cutoff 1 fragment_library_sort_method freq fragment_library_trans_origin no use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 ligand_atom_file ../001_structure/1s19_ligand_dockprep.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd no use_database_filter no orient_ligand yes automated_matching yes receptor_site_file ../002.surface_spheres/selected_spheres.sph max_orientations 1000 critical_points no chemical_matching no use_ligand_spheres no bump_filter no score_molecules no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.9_release/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.9_release/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.9_release/parameters/flex_drive.tbl ligand_outfile_prefix fragment.out write_orientations no num_scored_conformers 1 rank_ligands no
To run this, type:
dock6 -i fragment.in -o fragment.out
Once this has ran, the following files should have been generated: fraglib_linker.mol2, fraglib_rigid.mol2, fraglib_scaffold.mol2, fraglib_sidechain.mol2, fraglib_torenv.dat. To check the number of fragments in each file, you can type:
grep -wc MOLECULE *.mol2
You should get the following output upon using this command:
fraglib_linker.mol2:4 fraglib_rigid.mol2:0 fraglib_scaffold.mol2:2 fraglib_sidechain.mol2:2 fragment.out_scored.mol2:0
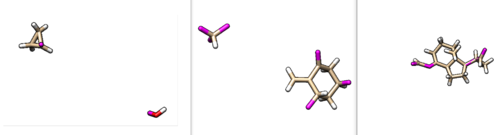
These fragments can also be visualized in Chimera. The results are shown below.
Focused Denovo Growth
We will be using the fragments that we have generated for the denovo dock. The different fragments will be connected together using restraints such as molecular weight and formal charges. DOCK will check the torsion environment file that was created in the fragment generation section to ensure reasonable connections are being made.
To begin, we will make a new directory and call it denovo_focus
mkdir denovo_focus