Rizzo Lab Research
PLEASE NOTE THIS PAGE IS BEING UPDATED
The Rizzo Research Group employ computational techniques to drug discovery projects. We are interested in both application and method development. We use two primary tools: docking and molecular dynamics (MD). Types of studies we perform include MD used to probe the origins of activity (free energy calculations), virtual screening for lead identification, and testset development to evaluate our methods. The current major focuses of the laboratory are outlined as follows.
Contents
HIV/AIDS
GP41 and viral membrane fusion inhibitor
| GP41 is a glycoprotein involved in viral membrane fusion. Our laboratory is interested in developing inhibitors that target GP41 and prevent the fusion event. To this end, we have constructed and all-atom model of T20 bound to gp41 and validated the model with all-atom molecular dynamics simulations. Virtual screening projects have also been performed to identify small molecule leads that target the hydrophobic pocket of gp41. Our collaborators have experimentally tested and identified molecules which exhibit strong activity.
McGillick, B. E.; Balius, T.E.; Mukherjee, S.; Rizzo, R. C. Origins of Resistance to the HIVgp41 Viral Entry Inhibitor T20. Biochemistry, 2010, 49 (17), 3575-3592 doi:10.1021/bi901915g PMID: 20230061 WEB PDF |
Cancer
EGFR and ErbB family
|
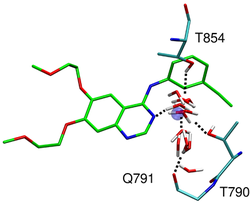
ErbB (epidermal growth factor receptor) family is a drug target for treating several types of cancers, including lung and breast cancers. ErbB family of receptor tyrosine kinases consists of EGFR, HER2, ErbB3, and ErbB4. Overexpression of EGFR is observed in 62% of NSCLC tumors(nonsmall cell lung cancer) and overexpression of EGFR and HER2 are important prognostic markers for breast cancer. Members of the ErbB family share a similar overall structural architecture comprising: (i) extracellular ligand binding domain, (ii) transmembrane domain, (iii) intracellular juxtamembrane domain, (iv) intracellular tyrosine kinase domain, and (v) C-terminal regulatory region where phosphorylation occurs. We are interested in targeting the tyrosine kinase domain(TKD). Approved small molecules of the TKD domain include erlotinib Tarceva, OSI Pharmaceuticals), gefitinib (Iressa, AstraZeneca), and lapatinib (Tykerb, Glaxo-SmithKline). A fourth compound called AEE788 (Novartis) is in development. Among them, erlotinib and gefitinib primarily target EGFR and lapatinib is a dual inhibitor of EGFR and ErbB2. Several cancer causing mutations or resistance mutations in EGFR and HER2 have been reported. We are interested in what is the driving force of binding and how these mutations affect binding. Through all-atom molecular dynamics simulations, water-mediated interactions seem to be especially important for understanding affinity and specificity for these systems.
|
Method development
DOCK development
The Rizzo Group co-develops the DOCK program [1] and contributed to the latest two releases:
The release v6.4, greatly improved the sampling behavior with the inclusion of internal energy during growth and minimization. See release notes.
The release v6.5 includes a new scoring function termed Footprint similarity score described below. See release notes.
Docking Testset Development
| To facilitate the improvements to the DOCK codebase and docking protocols, our group has developed a docking testset for pose reproduction.
See the following paper: Mukherjee, S.; Balius, T.E.; Rizzo, R. C. Docking Validation Resources: Protein Family and Ligand Flexibility Experiments. J. Chem. Inf. Model, 2010, 50, 1986-2000. PDF To obtain the testset visit Rizzo_Lab_Downloads. |
Chemical Sampling Method Development
|
De novo design algorithms are useful for both drug discovery and lead optimization. In de novo design, candidate molecules are assembled or grown from fragment libraries in the binding site of a protein target. Then, the affinity of the molecule can be predicted, typically through molecular mechanics-based scoring functions. Presumably, those molecules that are predicted to have higher affinity to the target protein would make better drug candidates. By building molecules from fragments in this way, one is not limited by the size of publicly-available virtual screening databases (ca. 106-107 molecules), which are exceedingly small when compared to the predicted size of actual chemical space (ca. 1065 molecules). However, de novo design can suffer from challenging obstacles including the inadvertent assembly of un-physical molecules, a combinatorial explosion in chemical space, and poor convergence. We have developed a novel de novo drug design method integrated into the infrastructure of the docking program DOCK6 which will be made avalible to the community in a future release. |
Docking Scoring Functions
|
Docking has two tasks: sampling and scoring. To Address scoring . . . Receptor flexibility is important for docking to address this . . . MultiGrid Docking |
|
|
Leveraging information from existing inhibitors is a useful paradigm for the discovery of new drugs. One scoring function developed in our group and implemented in DOCK 6.5, footprint similarity (FPS) score, uses an energetic profile or footprint of, for example, a known drug to identify a ligand which make similar interactions and is thus likely to bind. Other ideas for computationally generated references include molecular dynamics weighted ensembles, transition states and more. We are expanding this work on several levels including abstracting this scoring method to a grid-based method. See the following paper for more information on this topic. Balius, T.E.; Mukherjee, S.; Rizzo, R. C. Implementation and Evaluation of a Docking-Rescoring Method Using Molecular Footprint Comparisons. J. Comput. Chem., 2011, 32, 2273-2289. PDF |