2023 Denovo tutorial 1 with PDBID 4S0V
Contents
Introduction
DeNovo Design is where we are attempting to design a new ligand, usually with the hope of it binding more tightly to the protein, from scratch. There are three different ways of doing this:
- Generic DeNovo Design
- Focused Fragment Design
- DeNovo Refinement
This next tutorial will walk you through the details of each and is a continuation of the Virtual Screening tutorial. We will continue this work with #4s0v from the Protein Data Base.
Setting Up Your Environment
For this section we will need to create some more directories following this structure:
DeNovo Refinement
The DeNovo Refinement algorithm in Dock6.10 is an interesting way to determine the effects on a ligand/protein interaction by changing only part of the small molecule. The part of the ligand we want to experiment with is deleted from the structure, replaced with a dummy atom, and then run through DOCK. The program will try to find which residue can be placed in this now open position that will bind tightly to the protein.
Setting up the dummy atom
For the ligand from #4s0v, we will be removing a terminal ring and looking at what DOCK suggests to replace it with. The steps to do this are:
- Open the ligand minimized mol2 file we generated in the previous tutorial into Chimera.
- Open the protein into the same session
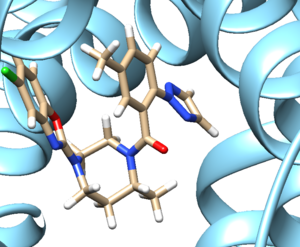
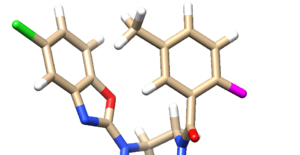
- Examine the binding site and choose a residue on the ligand that's pointing towards the inside of the binding site. For our protein this detailed section looks like:
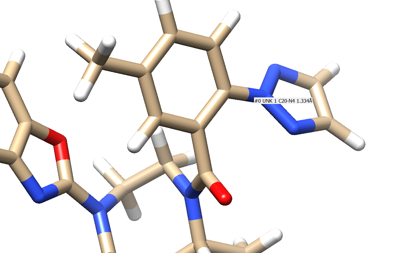
We see an imidazole ring pointing towards the binding site so will choose to work with that. Select the protein and hide it from view:
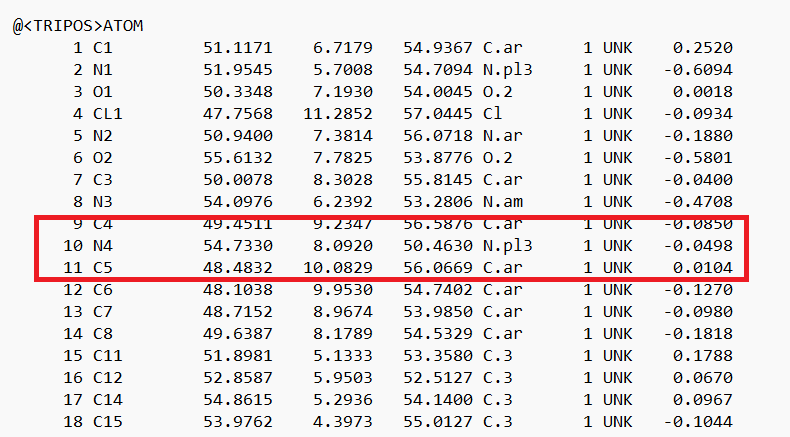
- Place your mouse over the atom connecting the ring to the rest of the ligand and note the atom and number. In this case it's N4.
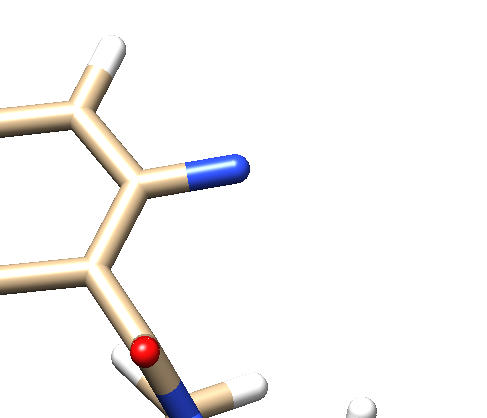
- Delete all the atoms from N4 to the end. Your ligand should now look something like:
- Save a .mol2 file of your ligand in this configuration. Make sure to give it a new filename such as 4s0v_denovoRefinement.mol2
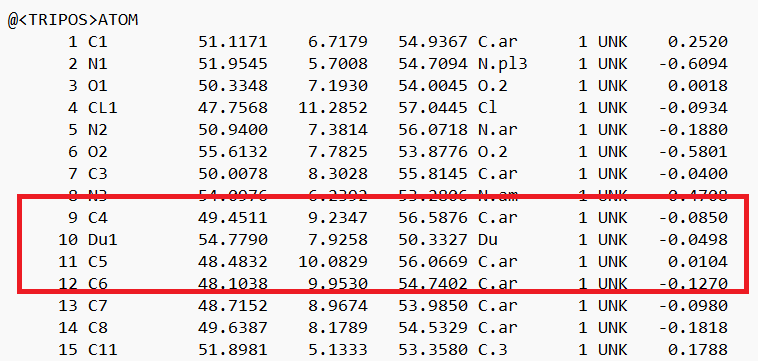
- Open the .mol2 file. If you're on a UNIX system, you can use vi; if you're on a PC, you can use textedit. Locate the atom that will now be changed to a dummy atom:
- Change the atom type to 'Du1' and the bond type to 'Du':
and save the file.
- Open a new session in Chimera and load the modified mol2 file. The "dummy" atom should now be purple:
- scp 4s0v_denovoRefinement.mol2 over to Seawulf and into the 012.denovoRefinement directory. From this point on we will be working on the command line.
Running DeNovo Refinement
Now that we have our .mol2 file on Seawulf we can run the DeNovo Refinement tool in DOCK6.10. We need to create an input file:
vi denovoRefinement.in
and type the following commands into it:
conformer_search_type denovo dn_fraglib_scaffold_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/fraglib_scaffold.mol2 dn_fraglib_linker_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/fraglib_linker.mol2 dn_fraglib_sidechain_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/fraglib_sidechain.mol2 dn_user_specified_anchor yes dn_fraglib_anchor_file 4s0v_ligand_denovo.mol2 dn_torenv_table /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/fraglib_torenv.dat dn_name_identifier 4s0v_denovo dn_sampling_method graph dn_graph_max_picks 30 dn_graph_breadth 3 dn_graph_depth 2 dn_graph_temperature 100.0 dn_pruning_conformer_score_cutoff 100.0 dn_pruning_conformer_score_scaling_factor 2.0 dn_pruning_clustering_cutoff 100.0 dn_mol_wt_cutoff_type soft dn_upper_constraint_mol_wt 1000.0 dn_lower_constraint_mol_wt 0.0 dn_mol_wt_std_dev 35.0 dn_constraint_rot_bon 15 dn_constraint_formal_charge 5 dn_heur_unmatched_num 1 dn_heur_matched_rmsd 2.0 dn_unique_anchors 1 dn_max_grow_layers 1 dn_max_root_size 25 dn_max_layer_size 25 dn_max_current_aps 5 dn_max_scaffolds_per_layer 1 dn_write_checkpoints yes dn_write_prune_dump no dn_write_orients no dn_write_growth_trees no dn_output_prefix 4s0v_denovo_output use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 use_database_filter no orient_ligand no bump_filter no score_molecules yes contact_score_primary no grid_score_primary yes grid_score_rep_rad_scale 1 grid_score_vdw_scale 1 grid_score_es_scale 1 grid_score_grid_prefix ../003.gridbox/grid minimize_ligand yes minimize_anchor no minimize_flexible_growth yes use_advanced_simplex_parameters no simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1 simplex_trans_step 1 simplex_rot_step 0.1 simplex_tors_step 10 simplex_grow_max_iterations 250 simplex_grow_tors_premin_iterations 0 simplex_random_seed 0 simplex_restraint_min yes simplex_coefficient_restraint 10 atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_de_novo.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl
To run your file simply type:
dock6 -i denovoRefinement.in -o denovoRefinement.out
Once the job has completed you will see the following new files in your directory:
- 2tio_denovo_output.anchor_1.root_layer_1.mol2
- 2tio_denovo_output.denovo_build.mol2
- denovoRefinement.out
Viewing New Molecules
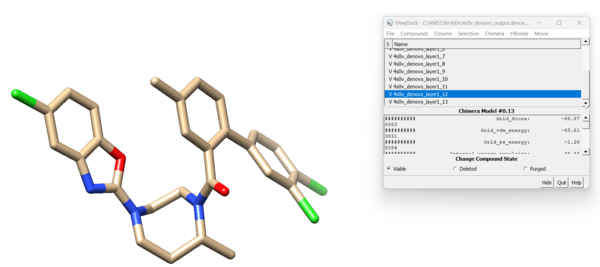
scp the two .mol2 files over to your local computer. Open a new session in Chimera. To view the new residues that DOCK added to the dummy node go to Tools → Surface/Binding Analysis → ViewDock and open 4s0v_denovo_output.denovo_build.mol2. As before, a dialogue box will open and you can click through the different model options.
Changing the input values in the input file can be powerful and gives us flexibility in how to "grow" our new ligand. These will be looked at more closely in the next two sections.
Focused DeNovo Design
The next two types of denovo design are similar to each other. We choose a structure, like an imidazole ring to "anchor" our ligand to the protein. DOCK will sample this structure all over the protein to find the best binding location. From that point a new ligand is "grown" out from this anchor point. The difference between focused and generic denovo design comes down to what residues DOCK uses to grow this new ligand.
Fragment Library Generation
In focused denovo design, a library of fragments which DOCK can use to grow the new ligand must be supplied to the program. We are "focusing" DOCK to work with specific fragments when building the new ligand. Very often when focused denovo design is used the fragment library is created using only the residues in the original ligand. This will be the approach we use for the following tutorial.
cd into your 013a.fragLib directory and create the input file:
vi fragLib.in
Type the following commands into the input file:
conformer_search_type flex write_fragment_libraries yes fragment_library_prefix 4s0v_fragLib fragment_library_freq_cutoff 1 fragment_library_sort_method freq fragment_library_trans_origin no use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 ligand_atom_file ../001.structure/4s0v_ligand_hydrogens.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd no use_database_filter no orient_ligand yes automated_matching yes receptor_site_file ../002.surface_spheres/selected_spheres.sph max_orientations 1000 critical_points no chemical_matching no use_ligand_spheres no bump_filter no score_molecules no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_de_novo.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl ligand_outfile_prefix 4s0v_focused write_orientations no num_scored_conformers 1 write_conformations no cluster_conformations yes cluster_rmsd_threshold 2.0 rank_ligands no
and run the program with:
dock6 -i fragLib.in -o fragLib.out
Once the program has successfully run you'll see the following new files in your directory:
- 4s0v_focused_scored.mol2
- 4s0v_fragLib_linker.mol2
- 4s0v_fragLib_rigid.mol2
- 4s0v_fragLib_scaffold.mol2
- 4s0v_fragLib_sidechain.mol2
- 4s0v_fragLib_torenv.dat
DeNovo Design
For this next section, cd into your 013b.focusGrowth directory.
There are three .mol2 files generated in the previous step that we need to give DOCK to perform the focused denovo growth. For this next step we again need an input file:
vi deNovoFocus.in
and type the following commands into the input file:
conformer_search_type denovo dn_fraglib_scaffold_file ../013a.fragLib/4s0v_fragLib_scaffold.mol2 dn_fraglib_linker_file ../013a.fragLib/4s0v_fragLib_linker.mol2 dn_fraglib_sidechain_file ../013a.fragLib/4s0v_fragLib_sidechain.mol2 dn_user_specified_anchor no dn_use_torenv_table yes dn_torenv_table ../013a.fragLib/4s0v_fragLib_torenv.dat dn_sampling_method graph dn_graph_max_picks 30 dn_graph_breadth 3 dn_graph_depth 2 dn_graph_temperature 100.0 dn_pruning_conformer_score_cutoff 100.0 dn_pruning_conformer_score_scaling_factor 1.0 dn_pruning_clustering_cutoff 100.0 dn_constraint_mol_wt 550.0 dn_constraint_rot_bon 15 dn_constraint_formal_charge 2.0 dn_heur_unmatched_num 1 dn_heur_matched_rmsd 2.0 dn_unique_anchors 2 dn_max_grow_layers 9 dn_max_root_size 25 dn_max_layer_size 25 dn_max_current_aps 5 dn_max_scaffolds_per_layer 1 dn_write_checkpoints yes dn_write_prune_dump no dn_write_orients no dn_write_growth_trees yes dn_output_prefix 4s0v_focused use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 use_database_filter no orient_ligand yes automated_matching yes receptor_site_file ../002.surface_spheres/selected_spheres.sph max_orientations 1000 critical_points no chemical_matching no use_ligand_spheres no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary yes grid_score_secondary no grid_score_rep_rad_scale 1 grid_score_vdw_scale 1 grid_score_es_scale 1 grid_score_grid_prefix ../003.gridbox/grid multigrid_score_secondary no dock3.5_score_secondary no continuous_score_secondary no footprint_similarity_score_secondary no pharmacophore_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand yes minimize_anchor yes minimize_flexible_growth yes use_advanced_simplex_parameters no simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1.0 simplex_trans_step 1.0 simplex_rot_step 0.1 simplex_tors_step 10.0 simplex_anchor_max_iterations 500 simplex_grow_max_iterations 500 simplex_grow_tors_premin_iterations 0 simplex_random_seed 0 simplex_restraint_min no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_de_novo.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl
Finally, to run this command type:
dock6 -i deNovoFocus.in -o deNovoFocus.out
Once this has successfully run you will see many new files in your directory. These all contain possible new ligands which can bind to your protein.
ReScoring Designed Molecules
The numerous molecules that were grown in the previous step need to be narrowed down to possible hits which can be investigated further. One way to do this is to reScore each ligand to find those that are similar to the original ligand in terms of interactions with the protein and overall composition. For these next steps, cd into your 013c.focusReScore directory and create your input file:
vi focusReScore.in
type the following commands into your input file:
conformer_search_type rigid use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 ligand_atom_file ../013b.focusGrowth/4s0v_focused.denovo_build.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd no use_database_filter no orient_ligand no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary no grid_score_secondary no multigrid_score_primary no multigrid_score_secondary no dock3.5_score_primary no dock3.5_score_secondary no continuous_score_primary no continuous_score_secondary no footprint_similarity_score_primary no footprint_similarity_score_secondary no pharmacophore_score_primary no pharmacophore_score_secondary no descriptor_score_primary yes descriptor_score_secondary no descriptor_use_grid_score no descriptor_use_multigrid_score no descriptor_use_continuous_score no descriptor_use_footprint_similarity yes descriptor_use_pharmacophore_score yes descriptor_use_tanimoto yes descriptor_use_hungarian yes descriptor_use_volume_overlap yes descriptor_fps_score_use_footprint_reference_mol2 yes descriptor_fps_score_footprint_reference_mol2_filename ../004.energy_min/4s0v.lig.min_scored.mol2 descriptor_fps_score_foot_compare_type Euclidean descriptor_fps_score_normalize_foot no descriptor_fps_score_foot_comp_all_residue yes descriptor_fps_score_receptor_filename ../001.structure/4s0v_protein_hydrogens_charges.mol2 descriptor_fps_score_vdw_att_exp 6 descriptor_fps_score_vdw_rep_exp 12 descriptor_fps_score_vdw_rep_rad_scale 1 descriptor_fps_score_use_distance_dependent_dielectric yes descriptor_fps_score_dielectric 4.0 descriptor_fps_score_vdw_fp_scale 1 descriptor_fps_score_es_fp_scale 1 descriptor_fps_score_hb_fp_scale 0 descriptor_fms_score_use_ref_mol2 yes descriptor_fms_score_ref_mol2_filename ../004.energy_min/4s0v.lig.min_scored.mol2 descriptor_fms_score_write_reference_pharmacophore_mol2 no descriptor_fms_score_write_reference_pharmacophore_txt no descriptor_fms_score_write_candidate_pharmacophore no descriptor_fms_score_write_matched_pharmacophore no descriptor_fms_score_compare_type overlap descriptor_fms_score_full_match yes descriptor_fms_score_match_rate_weight 5.0 descriptor_fms_score_match_dist_cutoff 1.0 descriptor_fms_score_match_proj_cutoff 0.7071 descriptor_fms_score_max_score 20 descriptor_fingerprint_ref_filename ../004.energy_min/4s0v.lig.min_scored.mol2 descriptor_hms_score_ref_filename ../004.energy_min/4s0v.lig.min_scored.mol2 descriptor_hms_score_matching_coeff -5 descriptor_hms_score_rmsd_coeff 1 descriptor_volume_score_reference_mol2_filename ../004.energy_min/4s0v.lig.min_scored.mol2 descriptor_volume_score_overlap_compute_method analytical descriptor_weight_fps_score 1 descriptor_weight_pharmacophore_score 1 descriptor_weight_fingerprint_tanimoto -1 descriptor_weight_hms_score 1 descriptor_weight_volume_overlap_score -1 gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_de_novo.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl chem_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/chem.defn pharmacophore_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/ph4.defn ligand_outfile_prefix 4s0v_focus_rescore write_footprints yes write_hbonds yes write_orientations no num_scored_conformers 1 rank_ligands no
Finally to run this command type:
dock6 -i focusReScore.in -o focusReScore.out
When the program has run successfully there will be three new files in your directory:
- 4s0v_focus_rescore_footprint_scored.txt
- 4s0v_focus_rescore_hbond_scored.txt
- 4s0v_focus_rescore_scored.mol2
To view these new ligands, scp the .mol2 file to your local computer and use ViewDock.
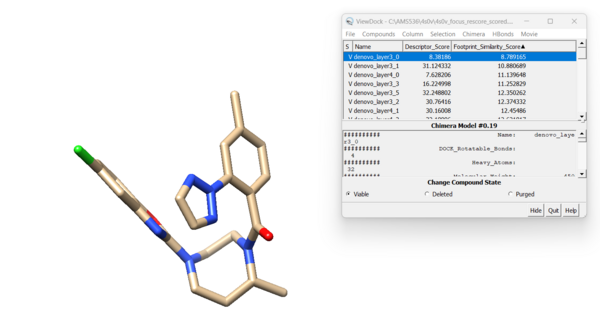
For this ligand we get 42 viable results. Looking at the new molecule with the lowest footprint score:
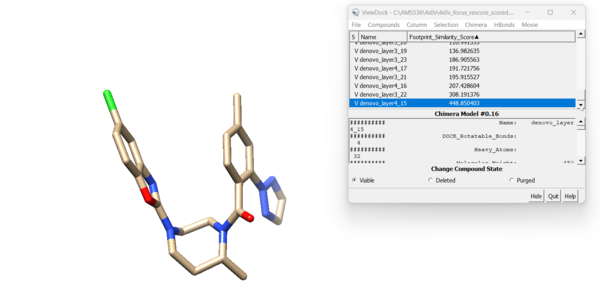
and the highest footprint score:
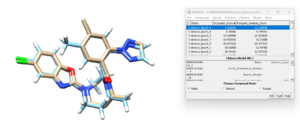
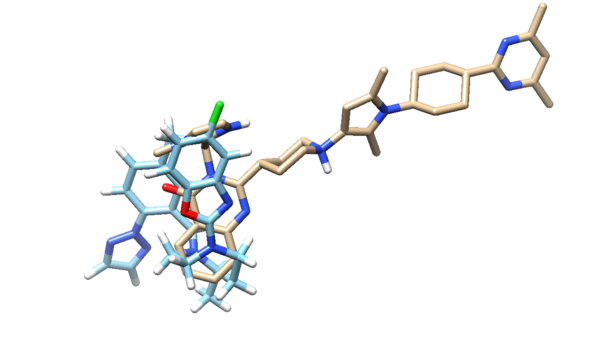
If we open the newly designed molecule with the lowest descriptor score and the energized minimized ligand from the previous tutorial, we see:
and we can see the two ligands are similar to each other although not exactly the same. This isn't too surprising since the library of fragments that DOCK had to work with originated from the residues in the original ligand. In the next section we give DOCK more freedom is choosing which residues to use in the denovo design.
Generic DeNovo Design
The final type of DeNovo design that we'll be looking at is Generic DeNovo Design. This is similar to the focused design in the previous section except we're not limiting DOCK to building the new molecule using only fragments present in the original ligand. This time we will be supplying DOCK with a library of many residues for it to use in generating the new molecule. This section will be done on the command line, so please cd to your 014.genericDenovo directory.
Create the input file:
vi genericLib.in
Type the following commands into the input file:
conformer_search_type denovo dn_fraglib_scaffold_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/fraglib_scaffold.mol2 dn_fraglib_linker_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/fraglib_linker.mol2 dn_fraglib_sidechain_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/fraglib_sidechain.mol2 dn_user_specified_anchor no dn_use_torenv_table yes dn_torenv_table /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/fraglib_torenv.dat dn_sampling_method graph dn_graph_max_picks 30 dn_graph_breadth 3 dn_graph_depth 2 dn_graph_temperature 100 dn_pruning_conformer_score_cutoff 100 dn_pruning_conformer_score_scaling_factor 1.0 dn_pruning_clustering_cutoff 100.0 dn_constraint_mol_wt 550.0 dn_constraint_rot_bon 15 dn_constraint_formal_charge 2.0 dn_heur_unmatched_num 1 dn_heur_matched_rmsd 2.0 dn_unique_anchors 1 dn_max_grow_layers 9 dn_max_root_size 25 dn_max_layer_size 25 dn_max_current_aps 5 dn_max_scaffolds_per_layer 1 dn_write_checkpoints yes dn_write_prune_dump no dn_write_orients no dn_write_growth_trees no dn_output_prefix 4s0v_genericDenovo_output use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 use_database_filter no orient_ligand yes automated_matching yes receptor_site_file ../002.surface_spheres/selected_spheres.sph max_orientations 1000 critical_points no chemical_matching no use_ligand_spheres no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary no grid_score_secondary no multigrid_score_primary no multigrid_score_secondary no dock3.5_score_primary no dock3.5_score_secondary no continuous_score_primary no continuous_score_secondary no footprint_similarity_score_primary no footprint_similarity_score_secondary no pharmacophore_score_primary no pharmacophore_score_secondary no descriptor_score_primary yes descriptor_score_secondary no descriptor_use_grid_score yes descriptor_use_pharmacophore_score no descriptor_use_tanimoto no descriptor_use_hungarian no descriptor_use_volume_overlap no descriptor_grid_score_rep_rad_scale 1 descriptor_grid_score_vdw_scale 1 descriptor_grid_score_es_scale 1 descriptor_grid_score_grid_prefix ../003.gridbox/grid descriptor_weight_grid_score 1 gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand yes minimize_anchor yes minimize_flexible_growth yes use_advanced_simplex_parameters no simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1.0 simplex_trans_step 1.0 simplex_rot_step 0.1 simplex_tors_step 10.0 simplex_anchor_max_iterations 500 simplex_grow_max_iterations 500 simplex_grow_tors_premin_iterations 0 simplex_random_seed 0 simplex_restraint_min no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_de_novo.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl
This calculation can be computationally expensive so it's best to run the input file with a slurm script:
#!/bin/bash # #SBATCH --job-name=4s0v_genericLib #SBATCH --output=genericLib_output.txt #SBATCH --ntasks-per-node=24 #SBATCH --nodes=1 #SBATCH --time=48:00:00 #SBATCH -p long-24core dock6 -i genericLib.in -o genericLib.out
Once the program is finished running you will see multiple new files in your directory including 4s0v_genericDenovo_output.denovo_build.mol2. scp this file over to your local computer. Start a new session in Chimera and open this file with ViewDock.
Using the command:
grep MOLECULE 4s0v_genericDenovo_output.denovo_build.mol2 | wc -l
you can determine how many new ligands DOCK "grew". In this case it's 537.
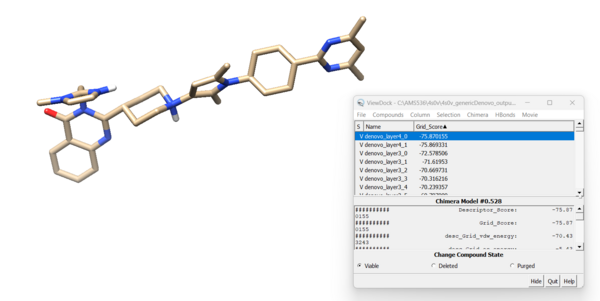
Looking at the molecule with the lowest grid score:
and if we overlay this molecule to the original minimized ligand:
we see that the ligands aren't the same at all. The original minimized ligand has a GridScore of -75.2 and this newly designed ligand has a GridScore of -75.8.
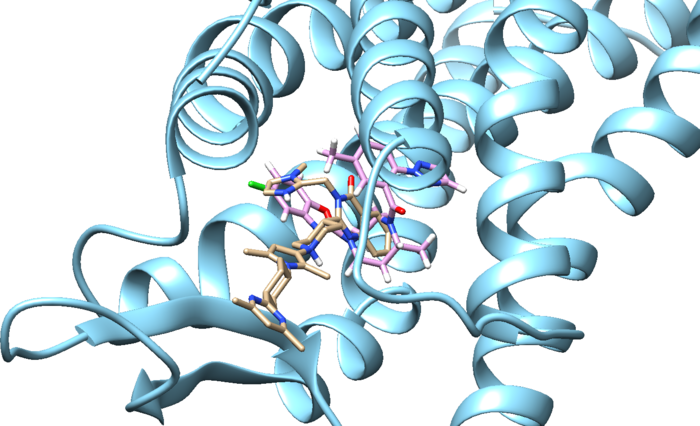
And finally we can look at both of these ligands interacting with the protein: