Difference between revisions of "2019 DOCK tutorial 1 with PDBID 2BXF"
Stonybrook (talk | contribs) (→Energy minimization) |
Stonybrook (talk | contribs) |
||
| (14 intermediate revisions by 2 users not shown) | |||
| Line 160: | Line 160: | ||
=V. Docking a single molecule for pose reproduction = | =V. Docking a single molecule for pose reproduction = | ||
| − | Under this section, the ligand for | + | Under this section, the ligand for 2BXF.pdb will be re-docked into the receptor. 3 Methods will be used to achieve this. |
1. rigid docking | 1. rigid docking | ||
| Line 185: | Line 185: | ||
read_mol_solvation no | read_mol_solvation no | ||
calculate_rmsd yes | calculate_rmsd yes | ||
| − | use_rmsd_reference_mol | + | use_rmsd_reference_mol yes |
| + | rmsd_reference_filename ../1.dockprep/2BXF_lig_withH.mol2 | ||
use_database_filter no | use_database_filter no | ||
orient_ligand no | orient_ligand no | ||
| Line 238: | Line 239: | ||
Run dock using the created input file. | Run dock using the created input file. | ||
| − | dock6 -i rigid.in | + | dock6 -i rigid.in -o rigid.out |
Follow a similar approach as we did for minimization to answer the prompted questions by either answering them manually using the answers in the lines below or by including the following lines in the input file before running dock. | Follow a similar approach as we did for minimization to answer the prompted questions by either answering them manually using the answers in the lines below or by including the following lines in the input file before running dock. | ||
| Line 244: | Line 245: | ||
conformer_search_type rigid | conformer_search_type rigid | ||
use_internal_energy yes | use_internal_energy yes | ||
| − | ligand_atom_file | + | ligand_atom_file 2BXF.lig.min_scored.mol2 |
limit_max_ligands no | limit_max_ligands no | ||
skip_molecule no | skip_molecule no | ||
read_mol_solvation no | read_mol_solvation no | ||
calculate_rmsd yes | calculate_rmsd yes | ||
| − | use_rmsd_reference_mol | + | use_rmsd_reference_mol yes |
| + | rmsd_reference_filename 2BXF.lig.min_scored.mol2 | ||
use_database_filter no | use_database_filter no | ||
orient_ligand yes | orient_ligand yes | ||
| Line 301: | Line 303: | ||
Open Chimera | Open Chimera | ||
| − | File -> Open -> | + | File -> Open -> 2BXF_rec_withH.mol2 |
| − | File -> Open -> | + | File -> Open -> 2BXF_lig_withH.mol2 |
Tools -> Surface/binding Analysis -> ViewDock -> Select the Rigid Dock output file. (rigid.out_scored.mol2) | Tools -> Surface/binding Analysis -> ViewDock -> Select the Rigid Dock output file. (rigid.out_scored.mol2) | ||
In the loaded dialog box select Dock4,5 or 6 | In the loaded dialog box select Dock4,5 or 6 | ||
| Line 314: | Line 316: | ||
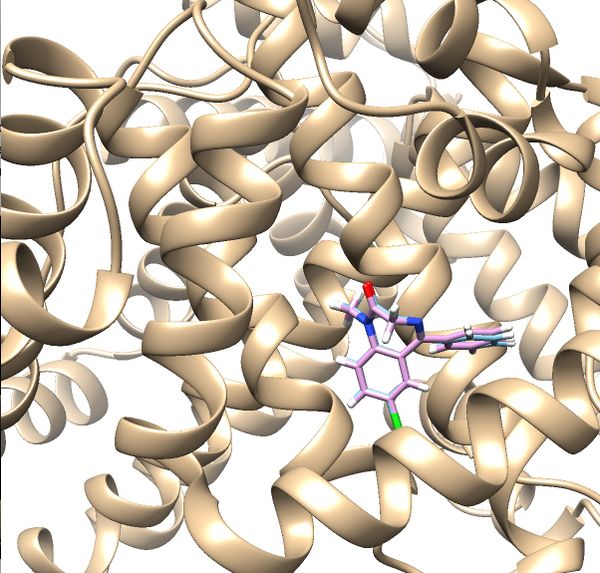
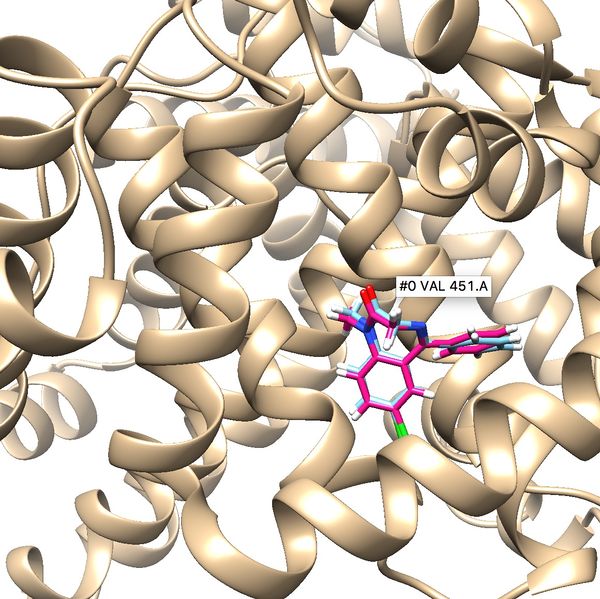
Your visualized structure should be similar to the image below. | Your visualized structure should be similar to the image below. | ||
| − | [[File: | + | [[File:2BXF rigiddocking.jpg|thumb|center|600px|Rigid docking results for 2BXF]] |
==Fixed Anchor Docking== | ==Fixed Anchor Docking== | ||
| Line 339: | Line 341: | ||
internal_energy_rep_exp 12 | internal_energy_rep_exp 12 | ||
internal_energy_cutoff 100.0 | internal_energy_cutoff 100.0 | ||
| − | ligand_atom_file ../1.dockprep/ | + | ligand_atom_file ../1.dockprep/2BXF_lig_withH.mol2 |
limit_max_ligands no | limit_max_ligands no | ||
skip_molecule no | skip_molecule no | ||
| Line 345: | Line 347: | ||
calculate_rmsd yes | calculate_rmsd yes | ||
use_rmsd_reference_mol yes | use_rmsd_reference_mol yes | ||
| − | rmsd_reference_filename ../1.dockprep/ | + | rmsd_reference_filename ../1.dockprep/2BXF_lig_withH.mol2 |
use_database_filter no | use_database_filter no | ||
orient_ligand no | orient_ligand no | ||
| Line 387: | Line 389: | ||
flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/flex.defn | flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/flex.defn | ||
flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/flex_drive.tbl | flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/flex_drive.tbl | ||
| − | ligand_outfile_prefix | + | ligand_outfile_prefix 2BXF_fad |
write_orientations no | write_orientations no | ||
num_scored_conformers 100 | num_scored_conformers 100 | ||
| Line 395: | Line 397: | ||
rank_ligands no | rank_ligands no | ||
| − | Once docking is completed an output file will be generated. ( | + | Once docking is completed an output file will be generated. (2BXF_fad_scored.mol2) Follow the same method used in rigid docking to visualize the docked poses using Chimera. Once it is visualized, it should like the image below. Notice all the poses 50 generated are in the same cluster and standard RMSD is 0.75. These indicate that docking is very successful. |
| − | [[File: | + | [[File:2BXF faddocking.jpg|thumb|center|600px| Poses generated for fixed anchor docking]] |
==Flexible Docking== | ==Flexible Docking== | ||
| Line 403: | Line 405: | ||
touch flex.in | touch flex.in | ||
Use the created input file to perform flexible docking using DOCK6. | Use the created input file to perform flexible docking using DOCK6. | ||
| − | dock6 -i flex.in | + | dock6 -i flex.in -o flex.out |
Answer the prompted questions using the following lines as we did in rigid and fixed anchor docking. | Answer the prompted questions using the following lines as we did in rigid and fixed anchor docking. | ||
| Line 422: | Line 424: | ||
internal_energy_rep_exp 12 | internal_energy_rep_exp 12 | ||
internal_energy_cutoff 100.0 | internal_energy_cutoff 100.0 | ||
| − | ligand_atom_file | + | ligand_atom_file 2BXF.lig.min_scored.mol2 |
limit_max_ligands no | limit_max_ligands no | ||
skip_molecule no | skip_molecule no | ||
read_mol_solvation no | read_mol_solvation no | ||
calculate_rmsd yes | calculate_rmsd yes | ||
| − | use_rmsd_reference_mol | + | use_rmsd_reference_mol yes |
| + | rmsd_reference_filename 2BXF.lig.min_scored.mol2 | ||
use_database_filter no | use_database_filter no | ||
orient_ligand yes | orient_ligand yes | ||
| Line 483: | Line 486: | ||
Once flexible docking is completed an output mol2 file will be generated. (flex.out_scored.mol2). Use the visualization steps used in rigid and fixed anchor docking and study the properties of the docking results. | Once flexible docking is completed an output mol2 file will be generated. (flex.out_scored.mol2). Use the visualization steps used in rigid and fixed anchor docking and study the properties of the docking results. | ||
| − | [[File: | + | [[File:2BXF_flexibledocking.jpg|thumb|center|600px| Flexible docking results for 2BXF]] |
==Molecular Footprint== | ==Molecular Footprint== | ||
| Line 498: | Line 501: | ||
conformer_search_type rigid | conformer_search_type rigid | ||
use_internal_energy no | use_internal_energy no | ||
| − | ligand_atom_file | + | ligand_atom_file 2BXF_lig_min.mol2 |
limit_max_ligands no | limit_max_ligands no | ||
skip_molecule no | skip_molecule no | ||
| Line 520: | Line 523: | ||
footprint_similarity_score_secondary no | footprint_similarity_score_secondary no | ||
fps_score_use_footprint_reference_mol2 yes | fps_score_use_footprint_reference_mol2 yes | ||
| − | fps_score_footprint_reference_mol2_filename | + | fps_score_footprint_reference_mol2_filename 2BXF_lig_with.mol2 |
fps_score_foot_compare_type Euclidean | fps_score_foot_compare_type Euclidean | ||
fps_score_normalize_foot no | fps_score_normalize_foot no | ||
fps_score_foot_comp_all_residue yes | fps_score_foot_comp_all_residue yes | ||
| − | fps_score_receptor_filename ../1.dockprep/ | + | fps_score_receptor_filename ../1.dockprep/2BXF_rec_withH.mol2 |
fps_score_vdw_att_exp 6 | fps_score_vdw_att_exp 6 | ||
fps_score_vdw_rep_exp 12 | fps_score_vdw_rep_exp 12 | ||
| Line 555: | Line 558: | ||
Use a python script to visualize the molecular footprint. The script can be accessed in the previous DOCK tutorials. Once the script is used, the molecular footprint should be similar to the image below. Notice the large deviations in energy at different amino acid residues. Those residues contribute more towards, the interaction between the ligand and therefore can be identified as important towards the binding of new ligands which can replace the original ligand in the PDB file. | Use a python script to visualize the molecular footprint. The script can be accessed in the previous DOCK tutorials. Once the script is used, the molecular footprint should be similar to the image below. Notice the large deviations in energy at different amino acid residues. Those residues contribute more towards, the interaction between the ligand and therefore can be identified as important towards the binding of new ligands which can replace the original ligand in the PDB file. | ||
| − | [[File: | + | [[File:2BXFfootprint.png|thumb|center|600px| Flexible docking results for 2BXF]] |
Latest revision as of 14:35, 9 March 2020
This tutorial contains a step by step approach to dock a known ligand to a known receptor.
Contents
I. Introduction
DOCK
DOCK is a molecular docking software that was originally developed at UCSF by Dr. Irwin Kuntz, Dr. Brian Shoicket, and colleagues. DOCK is used in drug discovery and molecular modeling. It consists of 2 main components: a search algorithm to explore chemical space for conformations and a scoring function to rank the results.
2BXF
The tutorial will be based on the PDB file 2BXF downloaded from the PDB Database. 2BXF is the crystal structure for human serum albumin complexed with diazepam.
Organization of Directories
We set up the files in our project space as such. It will be helpful to have these folders ready ahead of time.
0.files
1.dockprep
2.surface_spheres
3.gridbox
4.dock
6.footprint
7.virtual_screen
8.virtual_screen_mpi
9.cartesianmin
10.rescore
II. Preparation of the ligand and receptor
Download the pdb file 2BXF from Protein Data Bank and save the file in the 0.files folder.
Checking the Structure and Preparing the Complex without Hydrogens
- Open Chimera, open the downloaded pdb file, and check the structure for missing residues, gaps, heme groups, missing loops, and size. We recommend using proteins with no heme groups and proteins of a relatively smaller size. 2BXF did not have any of the preceding potential problems.
- Save the PDB file (the ligand and the receptor) as a mol2 file (2BXF_complex_noH.mol2) - When preparing both the ligand the receptor, you can open this complex mol2 file and delete the component that you won't need (eg. for preparing the receptor, open the complex, delete the ligand, and save).
Preparation of Receptor without Hydrogens
- Open 2BXF_complex_noH.mol2 through Chimera - Isolate the receptor by deleting the ligand. Click on the ligand and delete it (Actions -> Atoms/Bonds -> Delete). - Save the isolated receptor as a mol2 file (File -> Save mol2 -> 2BXF_rec_noH_mol2)
This receptor has been prepared without Hydrogens.
Preparation of Ligand without Hydrogens
- Open 2BXF_complex_noH.mol2 through Chimera again - Isolate the ligand by deleting the receptor. Click on the receptor and delete it (Actions -> Atoms/Bonds -> Delete). - Save the isolated ligand as a mol2 file (File -> Save mol2 -> 2BXF_lig_noH_mol2)
This ligand has been prepared without Hydrogens.
Preparation of receptor with Hydrogens
Preparation of ligand with Hydrogens
-Open the 2BXF_lig_noH.mol2 -Tools -> Surface Binding Analysis
Tools -> Structure Editing -> Add H (To add Hydrogen atoms)
Tools -> Structure Editing -> Add Charge (To add the charge use the latest AMBER force filed available for standard residues. Here we used AMBER ff14SB)
Save as a mol2 file. (2nnq_rec_withH.mol2)
- If you follow the step below all the above stated steps will automatically appear one after the other to prepare the receptor.
Tools -> Structure/Binding Analysis -> DockPrep
Preparation of ligand
- Open the PDB file via Chimera. - Using Chimera, isolate the ligand, add H atoms, add charge and save it as a mol2 file by following the same steps followed for the receptor.
Once all the files are prepared make sure to save the files in 1.dockprep folder.
III. Generating receptor surface and spheres
Preparation of DMS file
- Open 2BXF_rec_noH.mol2 using chimera. - Action -> Surface -> Show - Tools -> Structure Editing -> Write DMS - Save the 2BXF_rec_noH.dms into 3.surface_spheres folder
Reopen the file and make sure the surface was generated.
Transfer all the folders created so far to seawulf cluster to be used in DOCK.
Generating spheres
- Go to 2.surface_spheres folder - Create a new input file to create spheres by typing vim INSPH and type the following lines inside the file.
2BXF_rec_noH.dms R X 0.0 4.0 1.4 2BXF_rec.sph
The first line 2BXF_rec_noH.dms specifies the input file. R indicates that spheres generated will be outside of the receptor surface. X specifies all the points will be used. 0.0 is the distance in angstroms and it will avoid steric clashes. 4.0 is the maximum surface radius of the spheres and 1.4 is the minimum radius in angstroms.The last line 2BXF_spheres.sph creates the sph file that contains clustered spheres.
Once the INSPH file is ready, type the following command to generate the spheres.
sphgen -i INSPH -o OUTSPH
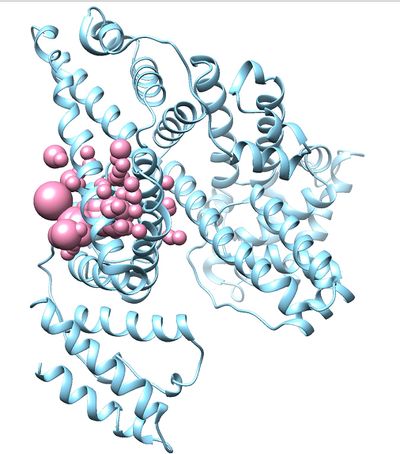
Once sphgen command is successful, 2BXF_spheres.sph file will be created. Open it up using Chimera along with 2BXF_rec_noH.mol2 file. You should get a similar output like the image below.
Selecting Spheres
Here we will be selecting the spheres which defines the binding pocket of the ligand because we are trying to direct the ligand towards that binding site rather than all over the receptor. To select the spheres type the following command.
sphere_selector 2BXF_rec.sph ../1.dockprep/2BXF_lig_withH.mol2 10.0
This command will select all of the spheres within 10.0 angstroms of the ligand and output them to selected_spheres.sph. Visualize the selected spheres using Chimera to make sure the correct spheres are selected. Notice that, spheres around the ligand binding site are kept and all the other spheres are deleted in the image below.
IV. Generating box and grid
Generating box
Move to 3.boxgrid directory Create a new file showbox.in and write the following lines in the file.
Y 8.0 ../2.surface_spheres/selected_spheres.sph 1 2BXF.box.pdb
Each of the above lines indicate that;
We intend to generate a box The box length should be 8 Angstroms Use the selected_spheres file in the designated location The name of the file that contains generated box.
Use the following command to generate the box.
showbox < showbox.in
If this step is successful, you should see a new file (2BXF.box.pdb) in 3.boxgrid folder.
Generating grid
Create a new file (grid.in)
Use the following command to generate the grid.
grid -i grid.in -o gridinfo.out
Answer the prompted questions with the answers given below. (or you can use the following lines and include them in the grid.in file before entering the above command. If you do that these questions won't be prompted again. They will be automatically answered by grid.in file created)
compute_grids yes grid_spacing 0.4 output_molecule no contact_score no energy_score yes energy_cutoff_distance 9999 atom_model a attractive_exponent 6 repulsive_exponent 9 distance_dielectric yes dielectric_factor 4 bump_filter yes bump_overlap 0.75 receptor_file ../1.dockprep/2BXF_rec_withH.mol2 box_file 2BXF.box.pdb vdw_definition_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/vdw_AMBER_parm99.defn score_grid_prefix grid
If the command is successful, three new files will be generated. (gridinfo.out, grid.nrg, grid.bmp). Go through gridinfo.out file to make sure all the information about the receptor in the file matches with the original information of the receptor. (Eg:- Total charge, residues and their charges) If the information doesn't match, that means you have made an error in one of the steps that you followed so far.
V. Docking a single molecule for pose reproduction
Under this section, the ligand for 2BXF.pdb will be re-docked into the receptor. 3 Methods will be used to achieve this.
1. rigid docking
2. fixed anchor docking
3. flexible docking
Energy minimization
Before performing docking, here the ligand will be subjected to energy minimization in order to remove unfavorable clashes. These clashes will affect rigid docking because in rigid docking the ligand will be docked as the complete ligand, whereas in other docking methods the ligand will be broken into fragments and the ligand will be built step by step considering favorable orientations and torsion angles after each fragment addition.
Go to the directory 4.dock and a create a new file (min.in) and enter the command below.
dock6 -i min.in -o min.out
Answer the prompted questions using the answers given below or include the following lines in the min.in file at before entering the above command to avoid answering the questions manually.
conformer_search_type rigid use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 ligand_atom_file ../1.dockprep/2BXF_lig_withH.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd yes use_rmsd_reference_mol yes rmsd_reference_filename ../1.dockprep/2BXF_lig_withH.mol2 use_database_filter no orient_ligand no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary yes grid_score_secondary no grid_score_rep_rad_scale 1 grid_score_vdw_scale 1 grid_score_es_scale 1 grid_score_grid_prefix ../3.boxgrid/grid multigrid_score_secondary no dock3.5_score_secondary no continuous_score_secondary no footprint_similarity_score_secondary no pharmacophore_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand yes simplex_max_iterations 1000 simplex_tors_premin_iterations 0 simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1.0 simplex_trans_step 1.0 simplex_rot_step 0.1 simplex_tors_step 10.0 simplex_random_seed 0 simplex_restraint_min yes simplex_coefficient_restraint 10.0 atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/flex_drive.tbl ligand_outfile_prefix 2BXF.lig.min write_orientations no num_scored_conformers 1 rank_ligands no
If the process is successful a new file (2BXF.lig.min_scored.mol2) will be generated. You can compare how is it changed from the initial structure by analyzing the RMSD value generated in the file. Visualize the new mol2 file along with receptor and the initial ligand mol2 files using Chimera to see the differences.
Rigid Docking
Create an input file for rigid docking
touch rigid.in
Run dock using the created input file.
dock6 -i rigid.in -o rigid.out
Follow a similar approach as we did for minimization to answer the prompted questions by either answering them manually using the answers in the lines below or by including the following lines in the input file before running dock.
conformer_search_type rigid use_internal_energy yes ligand_atom_file 2BXF.lig.min_scored.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd yes use_rmsd_reference_mol yes rmsd_reference_filename 2BXF.lig.min_scored.mol2 use_database_filter no orient_ligand yes automated_matching yes receptor_site_file ../2.surface_spheres/selected_spheres.sph max_orientations 1000 critical_points no chemical_matching no use_ligand_spheres no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary yes grid_score_secondary no grid_score_rep_rad_scale 1 grid_score_vdw_scale 1 grid_score_es_scale 1 grid_score_grid_prefix ../3.boxgrid/grid multigrid_score_secondary no dock3.5_score_secondary no continuous_score_secondary no footprint_similarity_score_secondary no pharmacophore_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand yes simplex_max_iterations 1000 simplex_tors_premin_iterations 0 simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1.0 simplex_trans_step 1.0 simplex_rot_step 0.1 simplex_tors_step 10.0 simplex_random_seed 0 simplex_restraint_min no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/flex_drive.tbl ligand_outfile_prefix rigid.out write_orientations no num_scored_conformers 1 rank_ligands no
Once rigid docking is successful, you will get an output file. (rigid.out_scored.mol2) Visualize the output file using Chimera by following steps to check the rigid docking success.
Open Chimera File -> Open -> 2BXF_rec_withH.mol2 File -> Open -> 2BXF_lig_withH.mol2 Tools -> Surface/binding Analysis -> ViewDock -> Select the Rigid Dock output file. (rigid.out_scored.mol2) In the loaded dialog box select Dock4,5 or 6
Once everything is loaded go to the ViewDock window and use it's menu to view all the calculated properties regarding the rigid docked ligand by following the steps below.
Column -> Show -> gridscore Column -> Show -> HA_RMSDs Follow the same steps to get all the properties
Your visualized structure should be similar to the image below.
Fixed Anchor Docking
Create an input file for fixed anchor docking.
touch fixed.in
Use the input file to perform fixed anchor docking
dock6 -i fixed.in
Use the following lines to answer the prompted questions as we did in rigid docking.
conformer_search_type flex user_specified_anchor no limit_max_anchors no min_anchor_size 5 pruning_use_clustering yes pruning_max_orients 1000 pruning_clustering_cutoff 100 pruning_conformer_score_cutoff 100.0 pruning_conformer_score_scaling_factor 1.0 use_clash_overlap no write_growth_tree no write_fragment_libraries no use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 ligand_atom_file ../1.dockprep/2BXF_lig_withH.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd yes use_rmsd_reference_mol yes rmsd_reference_filename ../1.dockprep/2BXF_lig_withH.mol2 use_database_filter no orient_ligand no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary yes grid_score_secondary no grid_score_rep_rad_scale 1 grid_score_vdw_scale 1 grid_score_es_scale 1 grid_score_grid_prefix ../2.boxgrid/grid multigrid_score_secondary no dock3.5_score_secondary no continuous_score_secondary no footprint_similarity_score_secondary no pharmacophore_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand yes minimize_anchor yes minimize_flexible_growth yes use_advanced_simplex_parameters no simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1.0 simplex_trans_step 1.0 simplex_rot_step 0.1 simplex_tors_step 10.0 simplex_anchor_max_iterations 500 simplex_grow_max_iterations 500 simplex_grow_tors_premin_iterations 0 simplex_random_seed 0 simplex_restraint_min no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/flex_drive.tbl ligand_outfile_prefix 2BXF_fad write_orientations no num_scored_conformers 100 write_conformations no cluster_conformations yes cluster_rmsd_threshold 2.0 rank_ligands no
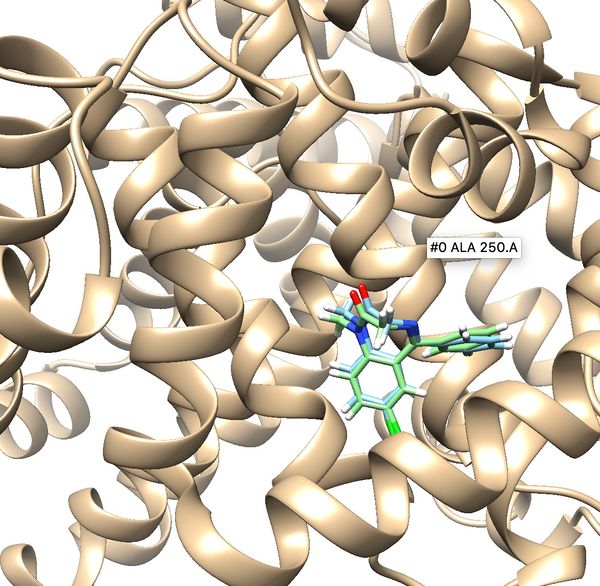
Once docking is completed an output file will be generated. (2BXF_fad_scored.mol2) Follow the same method used in rigid docking to visualize the docked poses using Chimera. Once it is visualized, it should like the image below. Notice all the poses 50 generated are in the same cluster and standard RMSD is 0.75. These indicate that docking is very successful.
Flexible Docking
Create a new input file for flexible docking. (flex.in)
touch flex.in
Use the created input file to perform flexible docking using DOCK6.
dock6 -i flex.in -o flex.out
Answer the prompted questions using the following lines as we did in rigid and fixed anchor docking.
conformer_search_type flex user_specified_anchor no limit_max_anchors no min_anchor_size 5 pruning_use_clustering yes pruning_max_orients 1000 pruning_clustering_cutoff 100 pruning_conformer_score_cutoff 100.0 pruning_conformer_score_scaling_factor 1.0 use_clash_overlap no write_growth_tree no write_fragment_libraries no use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 ligand_atom_file 2BXF.lig.min_scored.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd yes use_rmsd_reference_mol yes rmsd_reference_filename 2BXF.lig.min_scored.mol2 use_database_filter no orient_ligand yes automated_matching yes receptor_site_file ../2.surface_spheres/selected_spheres.sph max_orientations 1000 critical_points no chemical_matching no use_ligand_spheres no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary yes grid_score_secondary no grid_score_rep_rad_scale 1 grid_score_vdw_scale 1 grid_score_es_scale 1 grid_score_grid_prefix ../3.boxgrid/grid multigrid_score_secondary no dock3.5_score_secondary no continuous_score_secondary no footprint_similarity_score_secondary no pharmacophore_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand yes minimize_anchor yes minimize_flexible_growth yes use_advanced_simplex_parameters no simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1.0 simplex_trans_step 1.0 simplex_rot_step 0.1 simplex_tors_step 10.0 simplex_anchor_max_iterations 500 simplex_grow_max_iterations 500 simplex_grow_tors_premin_iterations 0 simplex_random_seed 0 simplex_restraint_min no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/flex_drive.tbl ligand_outfile_prefix flex.out write_orientations no num_scored_conformers 1 rank_ligands no
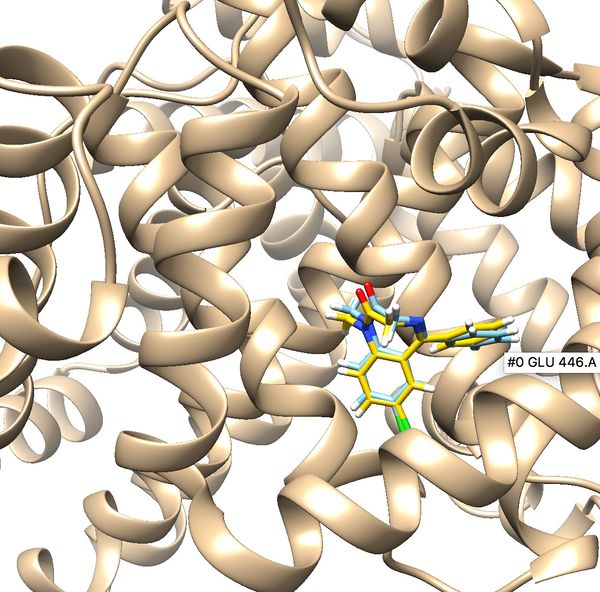
Once flexible docking is completed an output mol2 file will be generated. (flex.out_scored.mol2). Use the visualization steps used in rigid and fixed anchor docking and study the properties of the docking results.
Molecular Footprint
Molecular footprints can be used to determine how a ligand interacts with the receptor. Usually, the molecular footprint shows electrostatic interactions and Van der Waals interactions. Here, the molecular footprint will be used to determine how the ligand interacts with the receptor before and after minimization. To generate molecular footprints use following steps.
Go to directory 6.footprint
Generate an input file by typing;
touch footprint.in
Use DOCK6 to generate footprints
dock6 -i footprint.in
Use the following lines to answer the prompted questions.
conformer_search_type rigid use_internal_energy no ligand_atom_file 2BXF_lig_min.mol2 limit_max_ligands no skip_molecule no read_mol_solvation no calculate_rmsd no use_database_filter no orient_ligand no bump_filter no score_molecules yes contact_score_primary no contact_score_secondary no grid_score_primary no grid_score_secondary no multigrid_score_primary no multigrid_score_secondary no dock3.5_score_primary no dock3.5_score_secondary no continuous_score_primary no continuous_score_secondary no footprint_similarity_score_primary yes footprint_similarity_score_secondary no fps_score_use_footprint_reference_mol2 yes fps_score_footprint_reference_mol2_filename 2BXF_lig_with.mol2 fps_score_foot_compare_type Euclidean fps_score_normalize_foot no fps_score_foot_comp_all_residue yes fps_score_receptor_filename ../1.dockprep/2BXF_rec_withH.mol2 fps_score_vdw_att_exp 6 fps_score_vdw_rep_exp 12 fps_score_vdw_rep_rad_scale 1 fps_score_use_distance_dependent_dielectric yes fps_score_dielectric 4.0 fps_score_vdw_fp_scale 1 fps_score_es_fp_scale 1 fps_score_hb_fp_scale 0 pharmacophore_score_secondary no descriptor_score_secondary no gbsa_zou_score_secondary no gbsa_hawkins_score_secondary no SASA_score_secondary no amber_score_secondary no minimize_ligand no atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/vdw_AMBER_parm99.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6/parameters/flex_drive.tbl ligand_outfile_prefix footprint.out write_footprints yes write_hbonds yes write_orientations no num_scored_conformers 1 rank_ligands no
Once everything is successful these output files should be generated. (footprint.out_footprint_scored.txt, footprint.out_hbond_scored.txt, footprint.out_scored.mol2)
Use a python script to visualize the molecular footprint. The script can be accessed in the previous DOCK tutorials. Once the script is used, the molecular footprint should be similar to the image below. Notice the large deviations in energy at different amino acid residues. Those residues contribute more towards, the interaction between the ligand and therefore can be identified as important towards the binding of new ligands which can replace the original ligand in the PDB file.