2023 Denovo tutorial 2 with PDBID 3WZE
Contents
Introduction
This tutorial is a continuation of the virtual screening tutorial. In this tutorial, we'll continue to work with the receptor and ligand in PDB 3WZE, and we'll attempt to generate new ligands for the receptor using three kinds of de novo design: de novo refinement, focused de novo design, and generic de novo design.
De novo can be directly translated as "of new", but a more deft translation might be "from the beginning" or "from scratch". This method of ligand generation involves procedurally generating a ligand using algorithms within programs like DOCK, and is typically used to build entirely new ligands for proteins by building molecules outwards from an initial anchor one moiety at a time.
Generic de novo design best matches the prior description, in which a pre-selected or random anchor is positioned within the active site of the receptor, and then built outwards in a number of layers occupied by various sampled moieties. Focused de novo design is much like generic de novo design, except that the pool of sampled moieties is curtailed to suit the needs of the researcher. Finally, de novo refinement is when one begins with an already discovered ligand, then deletes some of the molecule and replaces it with a dummy atom, effectively using the remainder of the ligand as the anchor for the de novo design algorithms to modify.
Directories
Make new directories for de novo design:
mkdir 009_denovo_refine mkdir 010_denovo_focused mkdir 011_denovo_generic
De Novo Refinement
Ligand Preparation
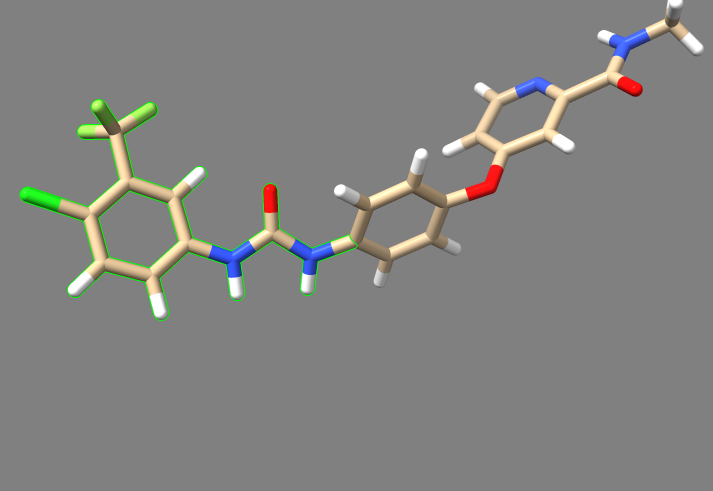
1. Open the final, energy minimized ligand mol2 file which was used for the 3WZE virtual screen tutorial, and also open the final receptor mol2 file that was used in that screen. Either Chimera or ChimeraX can be used to open the files. As long as no translations or rotations have occurred during the virtual screen process, the ligand should still be in its native orientation within the receptor's active site, as depicted by the original 3WZE pdb file.
2. Examine the binding pocket of the receptor, and choose a part of the 3WZE ligand that faces towards the interior of the binding pocket. Parts of the ligand that are innermost to the receptor make for the best parts to delete because they tend to have the most potential interactions with the protein, allowing the various groups tested in de novo design to have a better chance of interacting with a group on the protein. Choosing a part of the ligand to delete which faces the cytosol or the channel leading to the cytosol will be less likely to yield new ligands that can bind tightly to the interior of the receptor. To help recognize good sites for deletion, it's a good idea to show sidechains and hbonds, which can allow you to see which parts of the ligand are interacting with the protein.
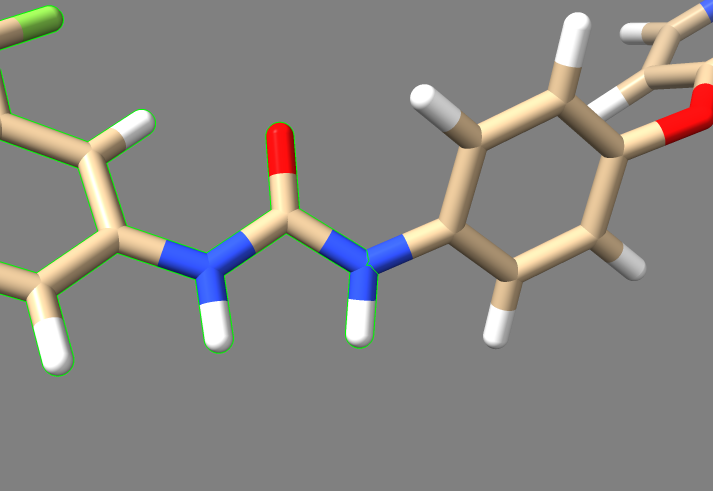
In this image, one can see the ligand sorafinib, and also the two hbonds that it forms with the nearby glutamic acid residue 71. It also forms an hbond with the backbone of the receptor using its amide oxygen. Based on this, we'll truncate those two amides and the entire aromatic ring closest to the camera. The camera is positioned to look from the side of the receptor where the binding pocket is deepest, so deleting everything closer than those amides will delete the parts of the ligand which are innermost.
3. Select and delete the receptor. Now that we've identified which part of the ligand to remove, we don't need the receptor anymore.
4. Orient the ligand so that the area you wish to delete is easy to see. Hold control down on your keyboard, then click and drag to cover the area. This should select the area.
5. Now deselect the first atom in the highlighted area. We're going to keep this atom so that it can be changed into a dummy atom. This style of de novo design requires a dummy atom to tell DOCK where to try putting new moieties, and it's easier to keep this nitrogen and change it into a dummy than it is to delete the whole selected area then manually attach a dummy.
6. Delete the selected area using Actions->Atoms/Bonds->Delete. Alternatively, if you're using ChimeraX, simply type "delete sel" into the command line.
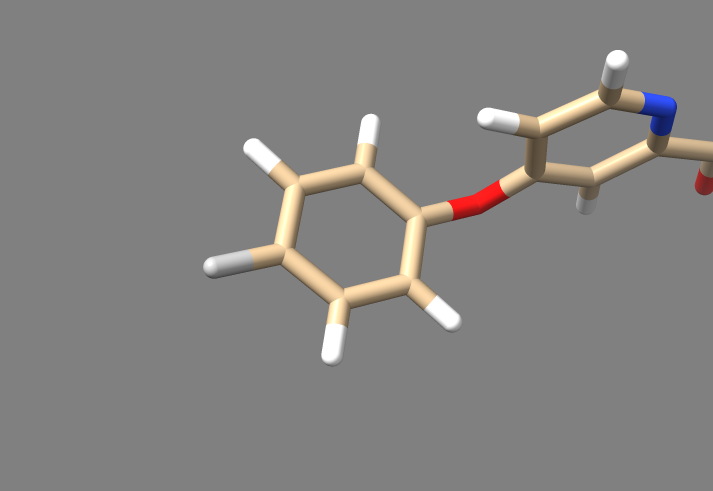
You should end up with a molecule that looks like this. Hover your mouse over that nitrogen we spared from deletion, and note its number. In this case the nitrogen is N14.
7. Save this truncated molecule as a mol2 file.
8. Open the mol2 file in a text editor on desktop, or with a command like "nano" from the command line.
9. Find N14, and change the atom type to "Du1". Also change its bond type to "Du".
10. To test whether the mol2 modification worked, open the mol2 with Chimera or ChimeraX. The dummy atom should appear purple or grey, respectively.
(Note that this image was taken with ChimeraX)
11. Now that our ligand is prepared, we can move it to Seawulf where we can perform the actual de novo refinement.
Running the Refinement
As with the virtual screen, DOCK can be run with an input file, the text of which will be shown below. However, it's a good idea to make your own input file rather than copying what is written here. That way, you can get a sense of what parameters can be adjusted before a de novo refinement run.
1. In the command line in Seawulf, type
touch de_novo_refine.in
Unlike "nano" or "vi", the "touch" command will allow you to make a blank file.
2. To go through the process of answering DOCK's many questions about your run, and to subsequently generate an input file, type
dock6 -i de_novo_refine.in
3. Answer the questions. Our input file is as follows:
conformer_search_type denovo dn_fraglib_scaffold_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/fraglib_scaffold.mol2 dn_fraglib_linker_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/fraglib_linker.mol2 dn_fraglib_sidechain_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/fraglib_sidechain.mol2 dn_user_specified_anchor yes dn_fraglib_anchor_file Chopped_ligand_for_denovo.mol2 dn_torenv_table /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/fraglib_torenv.dat dn_name_identifier 3WZE_refine dn_sampling_method graph dn_graph_max_picks 30 dn_graph_breadth 3 dn_graph_depth 2 dn_graph_temperature 100.0 dn_pruning_conformer_score_cutoff 100.0 dn_pruning_conformer_score_scaling_factor 2.0 dn_pruning_clustering_cutoff 100.0 dn_mol_wt_cutoff_type soft dn_upper_constraint_mol_wt 1000 dn_lower_constraint_mol_wt 0.0 dn_mol_wt_std_dev 35.0 dn_constraint_rot_bon 15 dn_constraint_formal_charge 5 dn_heur_unmatched_num 1 dn_heur_matched_rmsd 2.0 dn_unique_anchors 1 dn_max_grow_layers 1 dn_max_root_size 25 dn_max_layer_size 25 dn_max_current_aps 5 dn_max_scaffolds_per_layer 1 dn_write_checkpoints yes dn_write_prune_dump no dn_write_orients no dn_write_growth_trees no dn_output_prefix 3WZE_refine use_internal_energy yes internal_energy_rep_exp 12 internal_energy_cutoff 100.0 use_database_filter no orient_ligand no bump_filter no score_molecules yes contact_score_primary no grid_score_primary yes grid_score_rep_rad_scale 1 grid_score_vdw_scale 1 grid_score_es_scale 1 grid_score_grid_prefix grid minimize_ligand yes minimize_anchor no minimize_flexible_growth yes use_advanced_simplex_parameters no simplex_max_cycles 1 simplex_score_converge 0.1 simplex_cycle_converge 1.0 simplex_trans_step 1.0 simplex_rot_step 0.1 simplex_tors_step 10.0 simplex_grow_max_iterations 250 simplex_grow_tors_premin_iterations 0 simplex_random_seed 0 simplex_restraint_min yes simplex_coefficient_restraint 10.0 atom_model all vdw_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/vdw_de_novo.defn flex_defn_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex.defn flex_drive_file /gpfs/projects/AMS536/zzz.programs/dock6.10/parameters/flex_drive.tbl
4. Run the de novo refinenment in Seawulf using the following command:
dock6 -i de_novo_refine.in -o de_novo_refine.out
This should take a few minutes to complete, and when it does, there should be three new files:
3WZE_refine.anchor_1.root_layer_1.mol2 3WZE_refine.denovo_build.mol2 de_novo_refine.out
Checking the Results
1. Bring the two mol2 files to your local machine
2. If you're using Chimera, open Chimera and use Tools->Surface/Binding Analysis->ViewDock to open 3WZE_refine.denovo_build.mol2. If you're using ChimeraX, open that file, then type "viewdockx" into the command line.
Note that this image was taken from ChimeraX. Also, please disregard that I named my ligand "blooble".
3. Look through the results. Because we set "dn_max_grow_layers" to 1, this means that the dummy atom should only be replaced with a single fragment, and there should be no further fragments appended to the single one added.
For our results, we got 10 new molecules. If we had wanted more molecules, we could have increased the value of "dn_graph_max_picks" from 30 to a higher value. This means that dock would sample more fragments (We only got 10 molecules because 20 of the 30 picked fragments must have somehow been incompatible with the anchor).
If you want every possible new molecule based on the fragement library you're using, you could set "dn_sampling_method" to "exhaustive" instead of "graph".